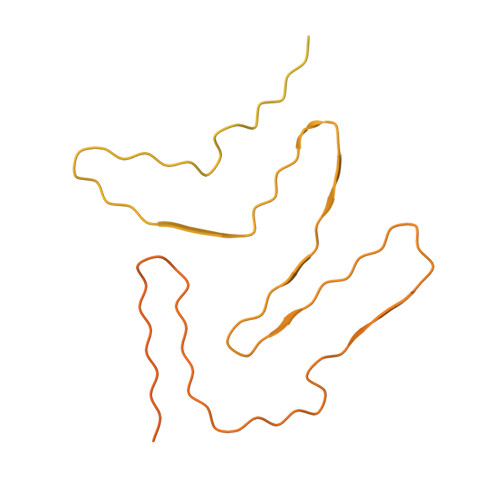
Molecular structure of an amyloid fibril formed by FUS low-complexity domain.
Sun, Y., Zhang, S., Hu, J., Tao, Y., Xia, W., Gu, J., Li, Y., Cao, Q., Li, D., Liu, C.(2022) iScience 25: 103701-103701
- PubMed: 35036880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2021.103701
- Primary Citation Related Structures:
7VQQ - PubMed Abstract:
FUS is a multifunctional nuclear protein which undergoes liquid-liquid phase separation in response to stress and DNA damage. Dysregulation of FUS dynamic phase separation leads to formation of pathological fibril closely associated with neurodegenerative diseases such as amyotrophic lateral sclerosis and frontotemporal dementia. In this study, we determined the cryo-EM structure of a cytotoxic fibril formed by the low-complexity (LC) domain of FUS at 2.9 Å resolution. The fibril structure exhibits a new and extensive serpentine fold consisting of three motifs incorporating together via a Tyr triad. FUS LC employs 91 residues to form an enlarged and stable fibril core via hydrophilic interaction and hydrogen bonds, which is distinct from most of previously determined fibrils commonly stabilized by hydrophobic interaction. Our work reveals the structural basis underlying formation of a cytotoxic and thermostable fibril of FUS LC and sheds light on understanding the liquid-to-solid phase transition of FUS in disease.
- Interdisciplinary Research Center on Biology and Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai 201210, China.
Organizational Affiliation: