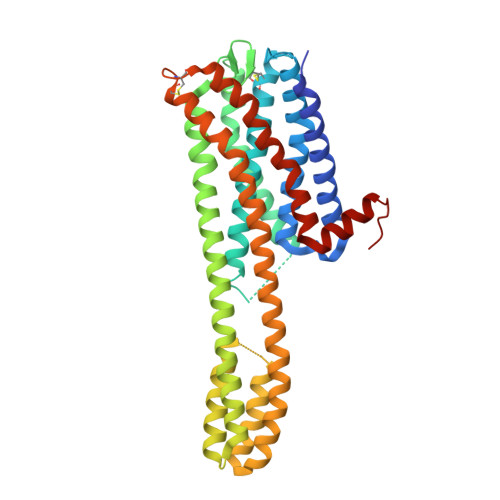
Structure-based design of a novel third-generation antipsychotic drug lead with potential antidepressant properties.
Chen, Z., Fan, L., Wang, H., Yu, J., Lu, D., Qi, J., Nie, F., Luo, Z., Liu, Z., Cheng, J., Wang, S.(2022) Nat Neurosci 25: 39-49
- PubMed: 34887590 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41593-021-00971-w
- Primary Citation Related Structures:
7VOD, 7VOE - PubMed Abstract:
Partial agonist activity at the dopamine D 2 receptor (DRD2) is a key feature of third-generation antipsychotics (TGAs). However, TGAs also act as antagonists or weak partial agonists to the serotonin (5-hydroxytryptamine; 5-HT) 2A receptor (5-HT 2A R). Here we present the crystal structures of aripiprazole- and cariprazine-bound human 5-HT 2A R. Both TGAs adopt an unexpected 'upside-down' pose in the 5-HT 2A R binding pocket, with secondary pharmacophores inserted in a similar way to a 'bolt'. This insight into the binding modes of TGAs offered a structural mechanism underlying their varied partial efficacies at 5-HT 2A R and DRD2. These structures enabled the design of a partial agonist at DRD2/3 and 5-HT 1A R with negligible 5-HT 2A R binding that displayed potent antipsychotic-like activity without motor side effects in mice. This TGA lead also had antidepressant-like effects and improved cognitive performance in mouse models via 5-HT 1A R. This work indicates that 5-HT 2A R affinity is a dispensable contributor to the therapeutic actions of TGAs.
- State Key Laboratory of Molecular Biology, Shanghai Institute of Biochemistry and Cell Biology, Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences, University of Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: