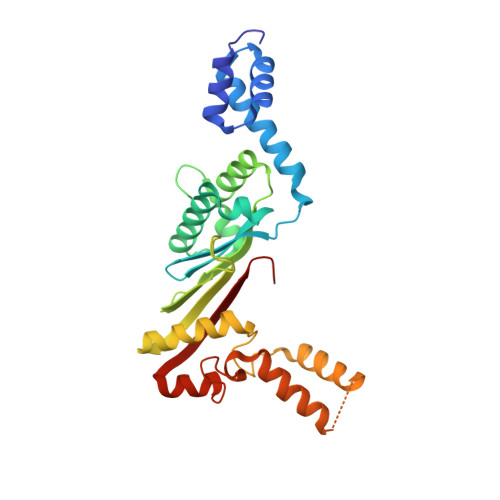
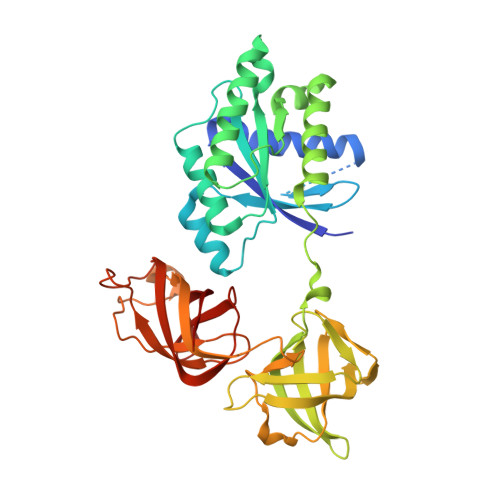
Structural insights of the elongation factor EF-Tu complexes in protein translation of Mycobacterium tuberculosis.
Zhan, B., Gao, Y., Gao, W., Li, Y., Li, Z., Qi, Q., Lan, X., Shen, H., Gan, J., Zhao, G., Li, J.(2022) Commun Biol 5: 1052-1052
- PubMed: 36192483 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-022-04019-y
- Primary Citation Related Structures:
7VMX, 7VOK - PubMed Abstract:
Tuberculosis (TB) caused by Mycobacterium tuberculosis (Mtb) is the second-deadliest infectious disease worldwide. Emerging evidence shows that the elongation factor EF-Tu could be an excellent target for treating Mtb infection. Here, we report the crystal structures of Mtb EF-Tu•EF-Ts and EF-Tu•GDP complexes, showing the molecular basis of EF-Tu's representative recycling and inactive forms in protein translation. Mtb EF-Tu binds with EF-Ts at a 1:1 ratio in solution and crystal packing. Mutation and SAXS analysis show that EF-Ts residues Arg13, Asn82, and His149 are indispensable for the EF-Tu/EF-Ts complex formation. The GDP binding pocket of EF-Tu dramatically changes conformations upon binding with EF-Ts, sharing a similar GDP-exchange mechanism in E. coli and T. ther. Also, the FDA-approved drug Osimertinib inhibits the growth of M. smegmatis, H37Ra, and M. bovis BCG strains by directly binding with EF-Tu. Thus, our work reveals the structural basis of Mtb EF-Tu in polypeptide synthesis and may provide a promising candidate for TB treatment.
- State Key Laboratory of Genetic Engineering, School of Life Sciences, Shanghai Engineering Research Center of Industrial Microorganisms, Fudan University, 200438, Shanghai, China.
Organizational Affiliation: