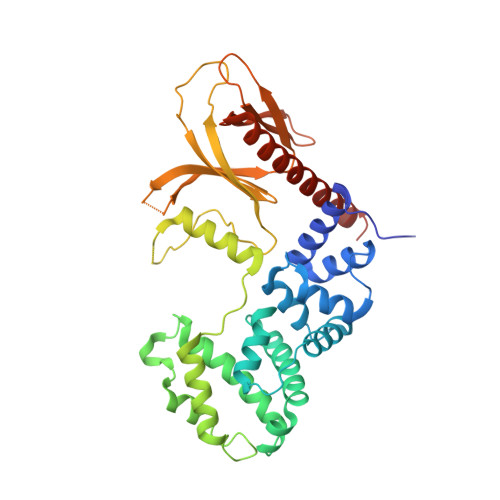
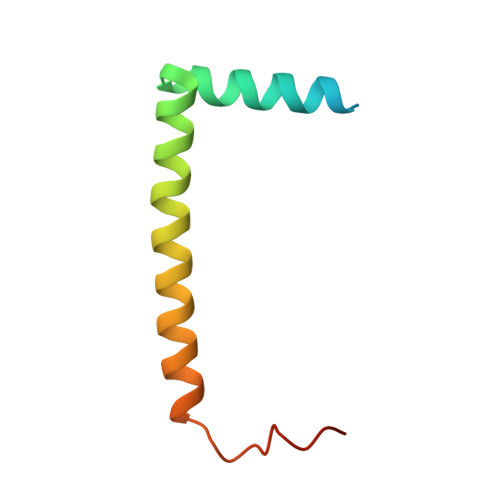
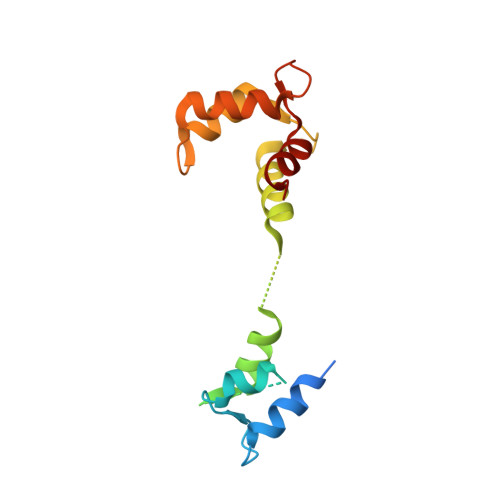
Ca2+-induced release of IQSEC2/BRAG1 autoinhibition under physiological and pathological conditions.
Bai, G., Li, H., Qin, P., Guo, Y., Yang, W., Lian, Y., Ye, F., Chen, J., Wu, M., Huang, R., Li, J., Lu, Y., Zhang, M.(2023) J Cell Biol 222
- PubMed: 37787765 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1083/jcb.202307117
- Primary Citation Related Structures:
7VMB - PubMed Abstract:
IQSEC2 (aka BRAG1) is a guanine nucleotide exchange factor (GEF) highly enriched in synapses. As a top neurodevelopmental disorder risk gene, numerous mutations are identified in Iqsec2 in patients with intellectual disabilities accompanied by other developmental, neurological, and psychiatric symptoms, though with poorly understood underlying molecular mechanisms. The atomic structures of IQSECs, together with biochemical analysis, presented in this study reveal an autoinhibition and Ca2+-dependent allosteric activation mechanism for all IQSECs and rationalize how each identified Iqsec2 mutation can alter the structure and function of the enzyme. Transgenic mice modeling two pathogenic variants of Iqsec2 (R359C and Q801P), with one activating and the other inhibiting the GEF activity of the enzyme, recapitulate distinct clinical phenotypes in patients. Our study demonstrates that different mutations on one gene such as Iqsec2 can have distinct neurological phenotypes and accordingly will require different therapeutic strategies.
- School of Life Sciences, Southern University of Science and Technology , Shenzhen, China.
Organizational Affiliation: