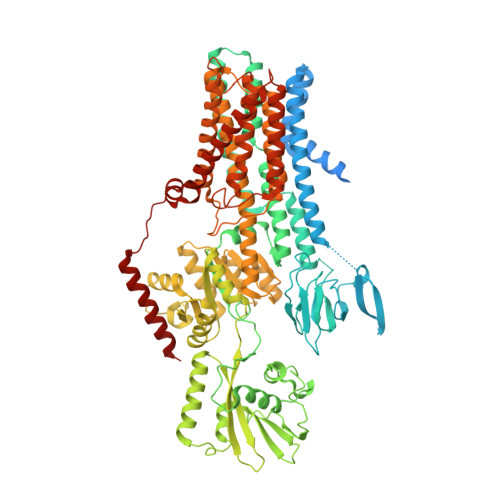
Structure and activation mechanism of the hexameric plasma membrane H + -ATPase.
Zhao, P., Zhao, C., Chen, D., Yun, C., Li, H., Bai, L.(2021) Nat Commun 12: 6439-6439
- PubMed: 34750373 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26782-y
- Primary Citation Related Structures:
7VH5, 7VH6 - PubMed Abstract:
The S. cerevisiae plasma membrane H + -ATPase, Pma1, is a P3A-type ATPase and the primary protein component of the membrane compartment of Pma1 (MCP). Like other plasma membrane H + -ATPases, Pma1 assembles and functions as a hexamer, a property unique to this subfamily among the larger family of P-type ATPases. It has been unclear how Pma1 organizes the yeast membrane into MCP microdomains, or why it is that Pma1 needs to assemble into a hexamer to establish the membrane electrochemical proton gradient. Here we report a high-resolution cryo-EM study of native Pma1 hexamers embedded in endogenous lipids. Remarkably, we found that the Pma1 hexamer encircles a liquid-crystalline membrane domain composed of 57 ordered lipid molecules. The Pma1-encircled lipid patch structure likely serves as the building block of the MCP. At pH 7.4, the carboxyl-terminal regulatory α-helix binds to the phosphorylation domains of two neighboring Pma1 subunits, locking the hexamer in the autoinhibited state. The regulatory helix becomes disordered at lower pH, leading to activation of the Pma1 hexamer. The activation process is accompanied by a 6.7 Å downward shift and a 40° rotation of transmembrane helices 1 and 2 that line the proton translocation path. The conformational changes have enabled us to propose a detailed mechanism for ATP-hydrolysis-driven proton pumping across the plasma membrane. Our structures will facilitate the development of antifungal drugs that target this essential protein.
- Department of Biochemistry and Biophysics, School of Basic Medical Sciences, Peking University, Beijing, China.
Organizational Affiliation: