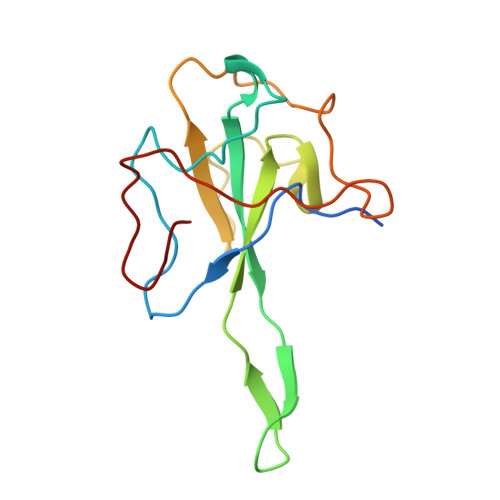
Atomic-Resolution Structure of SARS-CoV-2 Nucleocapsid Protein N-Terminal Domain.
Sarkar, S., Runge, B., Russell, R.W., Movellan, K.T., Calero, D., Zeinalilathori, S., Quinn, C.M., Lu, M., Calero, G., Gronenborn, A.M., Polenova, T.(2022) J Am Chem Soc 144: 10543-10555
- PubMed: 35638584 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.2c03320
- Primary Citation Related Structures:
7SD4, 7UW3 - PubMed Abstract:
The nucleocapsid (N) protein is one of the four structural proteins of the SARS-CoV-2 virus and plays a crucial role in viral genome organization and, hence, replication and pathogenicity. The N-terminal domain (N NTD ) binds to the genomic RNA and thus comprises a potential target for inhibitor and vaccine development. We determined the atomic-resolution structure of crystalline N NTD by integrating solid-state magic angle spinning (MAS) NMR and X-ray diffraction. Our combined approach provides atomic details of protein packing interfaces as well as information about flexible regions as the N- and C-termini and the functionally important RNA binding, β-hairpin loop. In addition, ultrafast (100 kHz) MAS 1 H-detected experiments permitted the assignment of side-chain proton chemical shifts not available by other means. The present structure offers guidance for designing therapeutic interventions against the SARS-CoV-2 infection.
- Department of Chemistry and Biochemistry, University of Delaware, Newark, Delaware 19716, United States.
Organizational Affiliation: