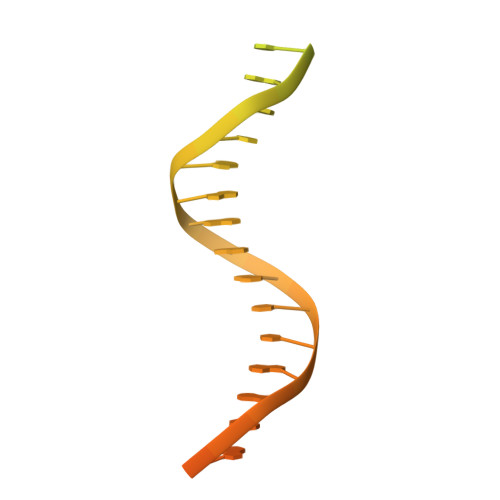
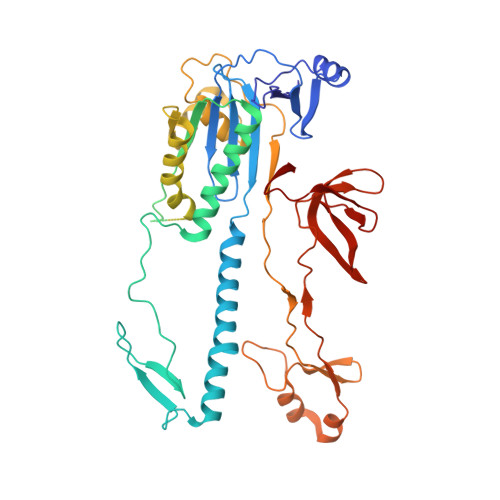
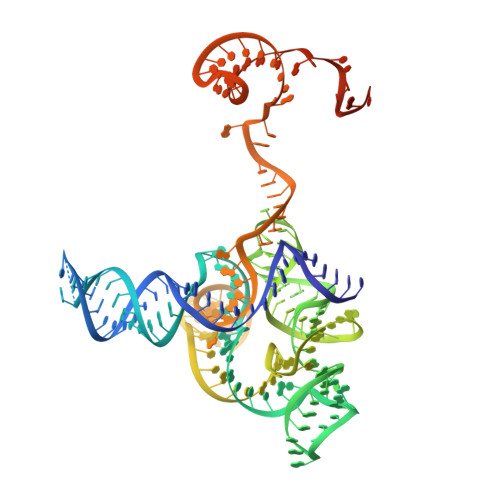
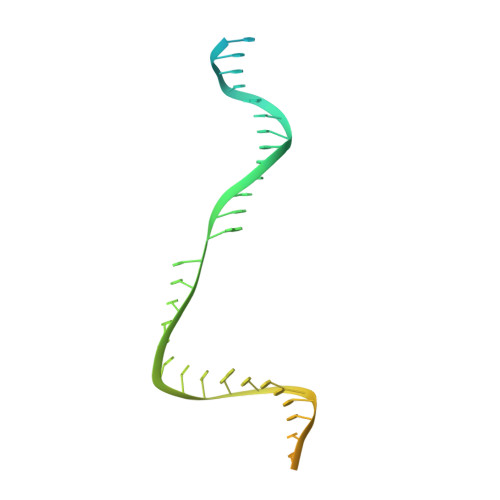
Structural basis for RNA-guided DNA cleavage by IscB-omega RNA and mechanistic comparison with Cas9.
Schuler, G., Hu, C., Ke, A.(2022) Science 376: 1476-1481
- PubMed: 35617371 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abq7220
- Primary Citation Related Structures:
7UTN, 8CSZ, 8CTL - PubMed Abstract:
Class 2 CRISPR effectors Cas9 and Cas12 may have evolved from nucleases in IS200/IS605 transposons. IscB is about two-fifths the size of Cas9 but shares a similar domain organization. The associated ωRNA plays the combined role of CRISPR RNA (crRNA) and trans-activating CRISPR RNA ( tracrRNA) to guide double-stranded DNA (dsDNA) cleavage. Here we report a 2.78-angstrom cryo-electron microscopy structure of IscB-ωRNA bound to a dsDNA target, revealing the architectural and mechanistic similarities between IscB and Cas9 ribonucleoproteins. Target-adjacent motif recognition, R-loop formation, and DNA cleavage mechanisms are explained at high resolution. ωRNA plays the equivalent function of REC domains in Cas9 and contacts the RNA-DNA heteroduplex. The IscB-specific PLMP domain is dispensable for RNA-guided DNA cleavage. The transition from ancestral IscB to Cas9 involved dwarfing the ωRNA and introducing protein domain replacements.
- Department of Molecular Biology and Genetics, Cornell University, 253 Biotechnology Building, Ithaca, NY 14853, USA.
Organizational Affiliation: