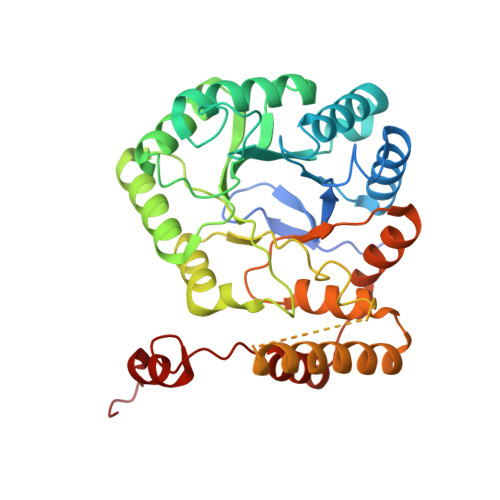
Structure-function characterization of an aldo-keto reductase involved in detoxification of the mycotoxin, deoxynivalenol.
Abraham, N., Schroeter, K.L., Zhu, Y., Chan, J., Evans, N., Kimber, M.S., Carere, J., Zhou, T., Seah, S.Y.K.(2022) Sci Rep 12: 14737-14737
- PubMed: 36042239 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-022-19040-8
- Primary Citation Related Structures:
7UTF - PubMed Abstract:
Deoxynivalenol (DON) is a mycotoxin, produced by filamentous fungi such as Fusarium graminearum, that causes significant yield losses of cereal grain crops worldwide. One of the most promising methods to detoxify this mycotoxin involves its enzymatic epimerization to 3-epi-DON. DepB plays a critical role in this process by reducing 3-keto-DON, an intermediate in the epimerization process, to 3-epi-DON. DepB Rleg from Rhizobium leguminosarum is a member of the new aldo-keto reductase family, AKR18, and it has the unusual ability to utilize both NADH and NADPH as coenzymes, albeit with a 40-fold higher catalytic efficiency with NADPH compared to NADH. Structural analysis of DepB Rleg revealed the putative roles of Lys-217, Arg-290, and Gln-294 in NADPH specificity. Replacement of these residues by site-specific mutagenesis to negatively charged amino acids compromised NADPH binding with minimal effects on NADH binding. The substrate-binding site of DepB Rleg is larger than its closest structural homolog, AKR6A2, likely contributing to its ability to utilize a wide range of aldehydes and ketones, including the mycotoxin, patulin, as substrates. The structure of DepB Rleg also suggests that 3-keto-DON can adopt two binding modes to facilitate 4-pro-R hydride transfer to either the re- or si-face of the C3 ketone providing a possible explanation for the enzyme's ability to convert 3-keto-DON to 3-epi-DON and DON in diastereomeric ratios of 67.2% and 32.8% respectively.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, Canada.
Organizational Affiliation: