Development of a GalNAc-Tyrosine-Specific Monoclonal Antibody and Detection of Tyrosine O -GalNAcylation in Numerous Human Tissues and Cell Lines.
Xia, L., Bellomo, T.R., Gibadullin, R., Congdon, M.D., Edmondson, E.F., Li, M., Wlodawer, A., Li, C., Temme, J.S., Patel, P., Butcher, D., Gildersleeve, J.C.(2022) J Am Chem Soc 144: 16410-16422
- PubMed: 36054098 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.2c04477
- Primary Citation Related Structures:
7UT3 - PubMed Abstract:
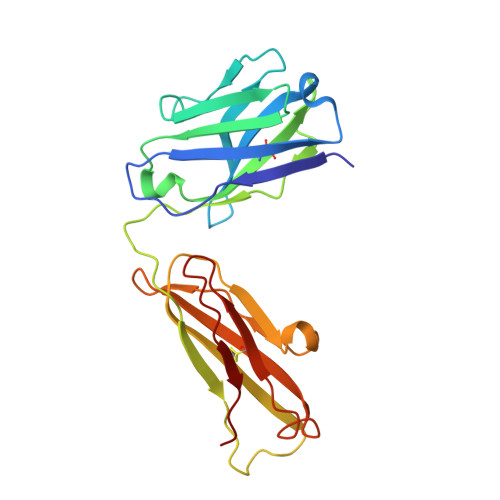
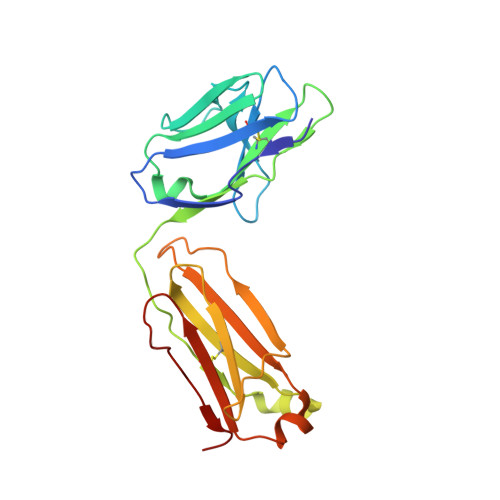
Glycosylation is a vital post-translational modification involved in a range of biological processes including protein folding, signaling, and cell-cell interactions. In 2011, a new type of O -linked glycosylation was discovered, wherein the side-chain oxygen of tyrosine is modified with a GalNAc residue (GalNAc-Tyr). At present, very little is known about GalNAc-Tyr prevalence, function, or biosynthesis. Herein, we describe the design and synthesis of a GalNAc-Tyr-derived hapten and its use in generating a GalNAc-Tyr selective monoclonal antibody. The antibody, G10C, has an unusually high affinity (app K D = 100 pM) and excellent selectivity for GalNAc-Tyr. We also obtained a crystal structure of the G10C Fab region in complex with 4-nitrophenyl- N -acetyl-α-d-galactosaminide (a small molecule mimic of GalNAc-Tyr) providing insights into the structural basis for high affinity and selectivity. Using this antibody, we discovered that GalNAc-Tyr is widely expressed in most human tissues, indicating that it is a ubiquitous and underappreciated post-translational modification. Localization to specific cell types and organ substructures within those tissues indicates that GalNAc-Tyr is likely regulated in a cell-specific manner. GalNAc-Tyr was also observed in a variety of cell lines and primary cells but was only present on the external cell surface in certain cancer cell lines, suggesting that GalNAc-Tyr localization may be altered in cancer cells. Collectively, the results shed new light on this under-studied form of glycosylation and provide access to new tools that will enable expanded biochemical and clinical investigations.
- Chemical Biology Laboratory, Center for Cancer Research, National Cancer Institute, Frederick, Maryland 21702, United States.
Organizational Affiliation: