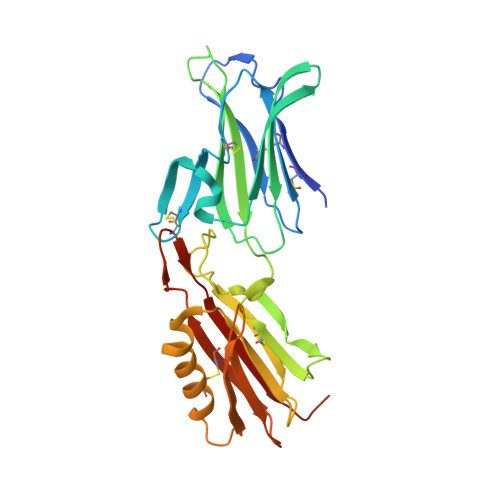
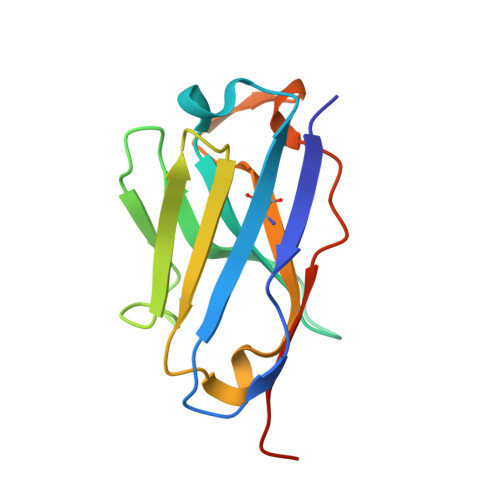
PCRCR complex is essential for invasion of human erythrocytes by Plasmodium falciparum.
Scally, S.W., Triglia, T., Evelyn, C., Seager, B.A., Pasternak, M., Lim, P.S., Healer, J., Geoghegan, N.D., Adair, A., Tham, W.H., Dagley, L.F., Rogers, K.L., Cowman, A.F.(2022) Nat Microbiol 7: 2039-2053
- PubMed: 36396942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-022-01261-2
- Primary Citation Related Structures:
7UNY, 7UNZ - PubMed Abstract:
The most severe form of malaria is caused by Plasmodium falciparum. These parasites invade human erythrocytes, and an essential step in this process involves the ligand PfRh5, which forms a complex with cysteine-rich protective antigen (CyRPA) and PfRh5-interacting protein (PfRipr) (RCR complex) and binds basigin on the host cell. We identified a heteromeric disulfide-linked complex consisting of P. falciparum Plasmodium thrombospondin-related apical merozoite protein (PfPTRAMP) and P. falciparum cysteine-rich small secreted protein (PfCSS) and have shown that it binds RCR to form a pentameric complex, PCRCR. Using P. falciparum lines with conditional knockouts, invasion inhibitory nanobodies to both PfPTRAMP and PfCSS, and lattice light-sheet microscopy, we show that they are essential for merozoite invasion. The PCRCR complex functions to anchor the contact between merozoite and erythrocyte membranes brought together by strong parasite deformations. We solved the structure of nanobody-PfCSS complexes to identify an inhibitory epitope. Our results define the function of the PCRCR complex and identify invasion neutralizing epitopes providing a roadmap for structure-guided development of these proteins for a blood stage malaria vaccine.
- The Walter and Eliza Hall Institute of Medical Research, Parkville, Australia.
Organizational Affiliation: