Metal-Mediated DNA Nanotechnology in 3D: Structural Library by Templated Diffraction.
Vecchioni, S., Lu, B., Livernois, W., Ohayon, Y.P., Yoder, J.B., Yang, C.F., Woloszyn, K., Bernfeld, W., Anantram, M.P., Canary, J.W., Hendrickson, W.A., Rothschild, L.J., Mao, C., Wind, S.J., Seeman, N.C., Sha, R.(2023) Adv Mater 35: e2210938-e2210938
- PubMed: 37268326 Search on PubMed
- DOI: https://doi.org/10.1002/adma.202210938
- Primary Citation Related Structures:
7UL8, 8E4D, 8E4E, 8EG4 - PubMed Abstract:
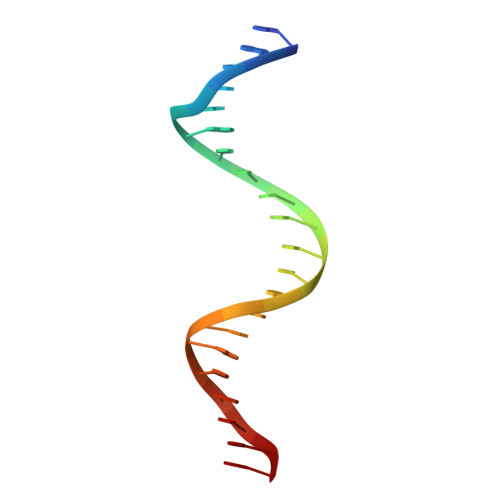
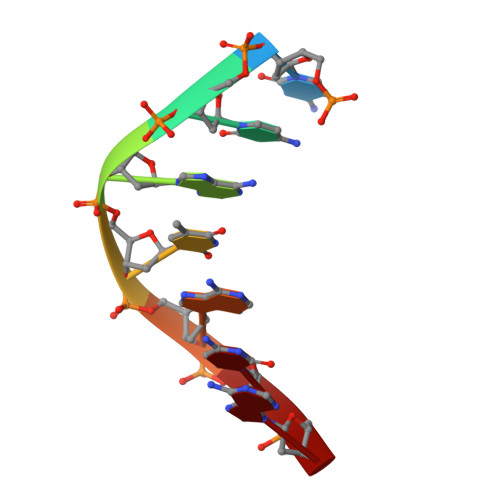
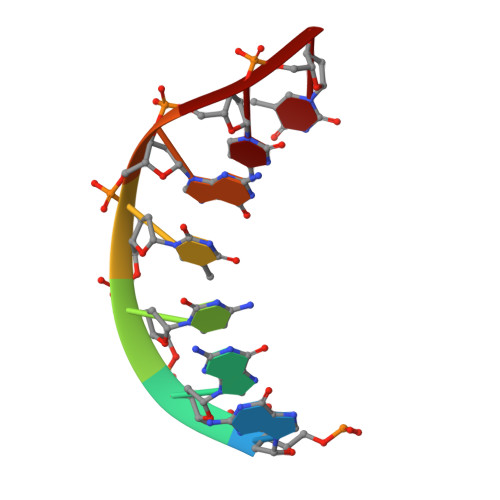
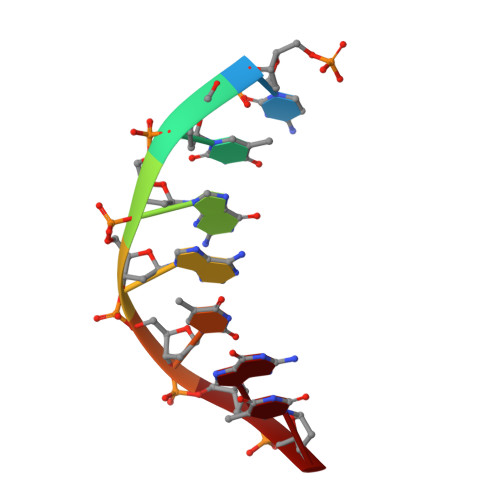
DNA double helices containing metal-mediated DNA (mmDNA) base pairs are constructed from Ag + and Hg 2+ ions between pyrimidine:pyrimidine pairs with the promise of nanoelectronics. Rational design of mmDNA nanomaterials is impractical without a complete lexical and structural description. Here, the programmability of structural DNA nanotechnology toward its founding mission of self-assembling a diffraction platform for biomolecular structure determination is explored. The tensegrity triangle is employed to build a comprehensive structural library of mmDNA pairs via X-ray diffraction and generalized design rules for mmDNA construction are elucidated. Two binding modes are uncovered: N3-dominant, centrosymmetric pairs and major groove binders driven by 5-position ring modifications. Energy gap calculations show additional levels in the lowest unoccupied molecular orbitals (LUMO) of mmDNA structures, rendering them attractive molecular electronic candidates.
- Department of Chemistry, New York University, New York, NY, 10003, USA.
Organizational Affiliation: