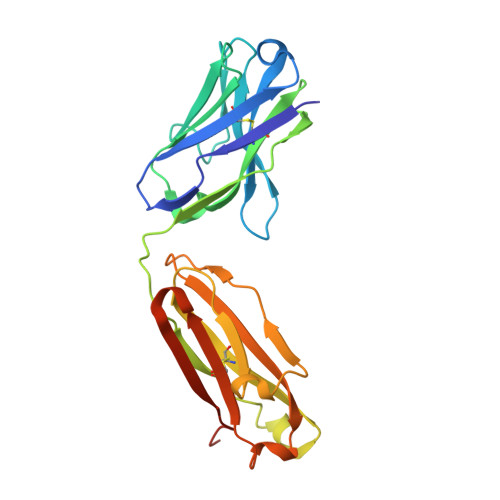
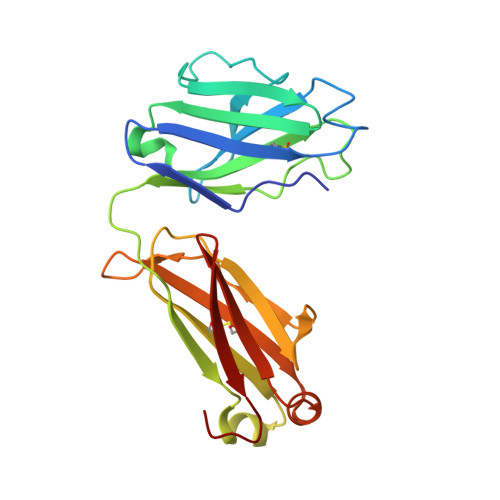
Genetic and structural basis of the human anti-alpha-galactosyl antibody response.
Langley, D.B., Schofield, P., Nevoltris, D., Jackson, J., Jackson, K.J.L., Peters, T.J., Burk, M., Matthews, J.M., Basten, A., Goodnow, C.C., van Nunen, S., Reed, J.H., Christ, D.(2022) Proc Natl Acad Sci U S A 119: e2123212119-e2123212119
- PubMed: 35867757 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2123212119
- Primary Citation Related Structures:
7UEL, 7UEM, 7UEN - PubMed Abstract:
Humans lack the capacity to produce the Galα1-3Galβ1-4GlcNAc (α-gal) glycan, and produce anti-α-gal antibodies upon exposure to the carbohydrate on a diverse set of immunogens, including commensal gut bacteria, malaria parasites, cetuximab, and tick proteins. Here we use X-ray crystallographic analysis of antibodies from α-gal knockout mice and humans in complex with the glycan to reveal a common binding motif, centered on a germline-encoded tryptophan residue at Kabat position 33 (W33) of the complementarity-determining region of the variable heavy chain (CDRH1). Immunoglobulin sequencing of anti-α-gal B cells in healthy humans and tick-induced mammalian meat anaphylaxis patients revealed preferential use of heavy chain germline IGHV3-7, encoding W33, among an otherwise highly polyclonal antibody response. Antigen binding was critically dependent on the presence of the germline-encoded W33 residue for all of the analyzed antibodies; moreover, introduction of the W33 motif into naive IGHV3-23 antibody phage libraries enabled the rapid selection of α-gal binders. Our results outline structural and genetic factors that shape the human anti-α-galactosyl antibody response, and provide a framework for future therapeutics development.
- Garvan Institute of Medical Research, Darlinghurst, NSW 2010, Australia.
Organizational Affiliation: