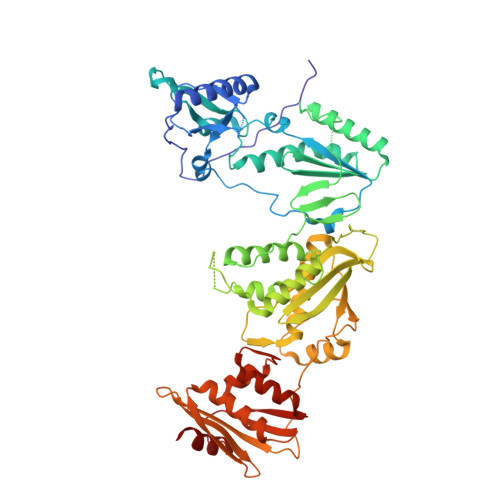
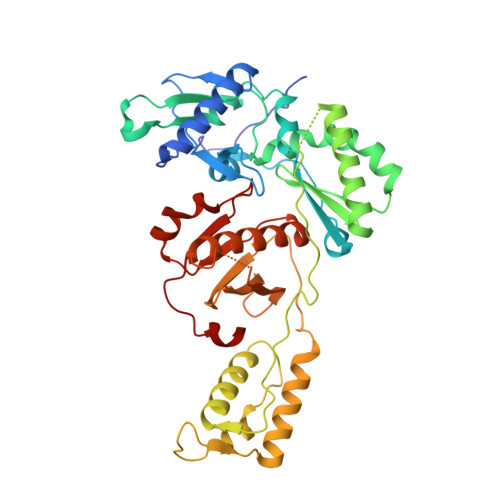
Design, synthesis, and biological testing of biphenylmethyloxazole inhibitors targeting HIV-1 reverse transcriptase.
Carter, Z.J., Hollander, K., Spasov, K.A., Anderson, K.S., Jorgensen, W.L.(2023) Bioorg Med Chem Lett 84: 129216-129216
- PubMed: 36871704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2023.129216
- Primary Citation Related Structures:
7U5Z - PubMed Abstract:
We report non-nucleoside inhibitors of HIV-1 reverse transcriptase (NNRTIs) using a biphenylmethyloxazole pharmacophore. A crystal structure of benzyloxazole 1 was obtained and suggested the potential viability of biphenyl analogues. In particular, 6a, 6b, and 7 turned out to be potent NNRTIs with low-nanomolar activity in enzyme inhibition and infected T-cell assays, and with low cytotoxicity. Though modeling further suggested that analogues with fluorosulfate and epoxide warheads might provide covalent modification of Tyr188, synthesis and testing did not find evidence for this outcome.
- Department of Chemistry, Yale University, New Haven, CT 06520-8107, USA.
Organizational Affiliation: