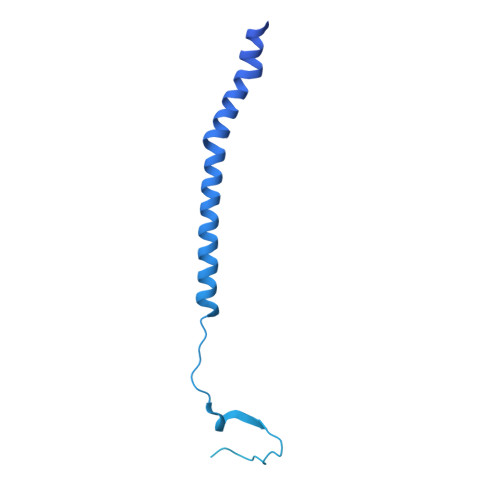
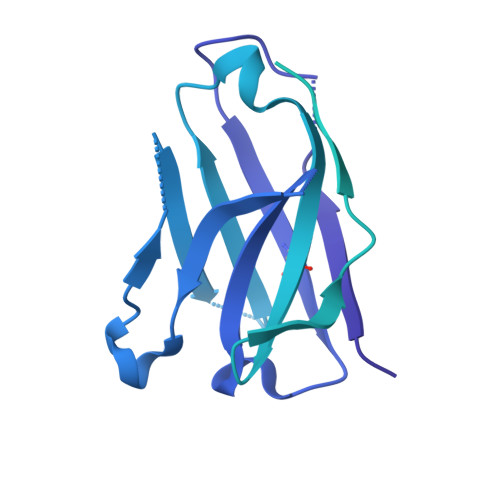
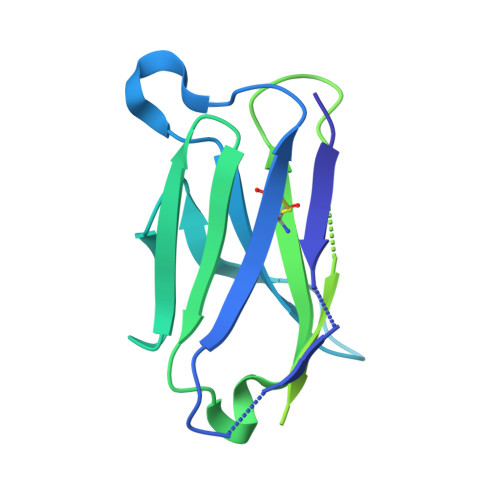
Architecture and antigenicity of the Nipah virus attachment glycoprotein.
Wang, Z., Amaya, M., Addetia, A., Dang, H.V., Reggiano, G., Yan, L., Hickey, A.C., DiMaio, F., Broder, C.C., Veesler, D.(2022) Science 375: 1373-1378
- PubMed: 35239409 Search on PubMed
- DOI: https://doi.org/10.1126/science.abm5561
- Primary Citation Related Structures:
7TXZ, 7TY0 - PubMed Abstract:
Nipah virus (NiV) and Hendra virus (HeV) are zoonotic henipaviruses (HNVs) responsible for outbreaks of encephalitis and respiratory illness. The entry of HNVs into host cells requires the attachment (G) and fusion (F) glycoproteins, which are the main targets of antibody responses. To understand viral infection and host immunity, we determined a cryo-electron microscopy structure of the NiV G homotetrameric ectodomain in complex with the nAH1.3 broadly neutralizing antibody Fab fragment. We show that a cocktail of two nonoverlapping G-specific antibodies neutralizes NiV and HeV synergistically and limits the emergence of escape mutants. Analysis of polyclonal serum antibody responses elicited by vaccination of macaques with NiV G indicates that the receptor binding head domain is immunodominant. These results pave the way for implementing multipronged therapeutic strategies against these deadly pathogens.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: