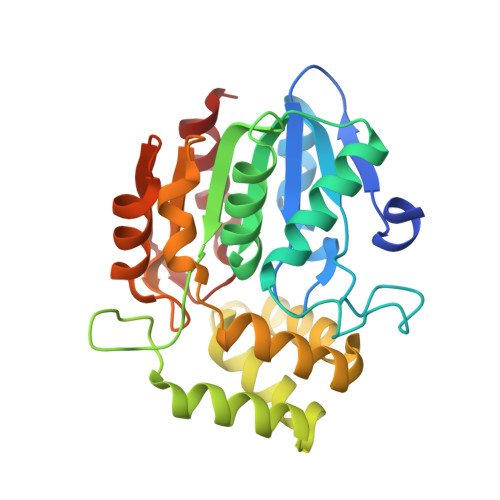
Crystal structure of Arabidopsis DWARF14-LIKE2 (DLK2) reveals a distinct substrate binding pocket architecture.
Burger, M., Honda, K., Kondoh, Y., Hong, S., Watanabe, N., Osada, H., Chory, J.(2022) Plant Direct 6: e446-e446
- PubMed: 36172078 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pld3.446
- Primary Citation Related Structures:
7TVW - PubMed Abstract:
In Arabidopsis thaliana , the Sigma factor B regulator RsbQ-like family of α/β hydrolases contains the strigolactone (SL) receptor DWARF14 (AtD14), the karrikin receptor KARRIKIN INSENSITIVE2 (AtKAI2), and DWARF14-LIKE2 (AtDLK2), a protein of unknown function. Despite very similar protein folds, AtD14 and AtKAI2 differ in size and architecture of their ligand binding pockets, influencing their substrate specificity. We present the 1.5 Å crystal structure of AtDLK2, revealing the smallest ligand binding pocket in the protein family, bordered by two unique glycine residues. We identified a gatekeeper residue in the protein's lid domain and present a pyrrolo-quinoline-dione compound that inhibits AtDLK2's enzymatic activity.
- Plant Biology Laboratory Salk Institute for Biological Studies La Jolla California USA.
Organizational Affiliation: