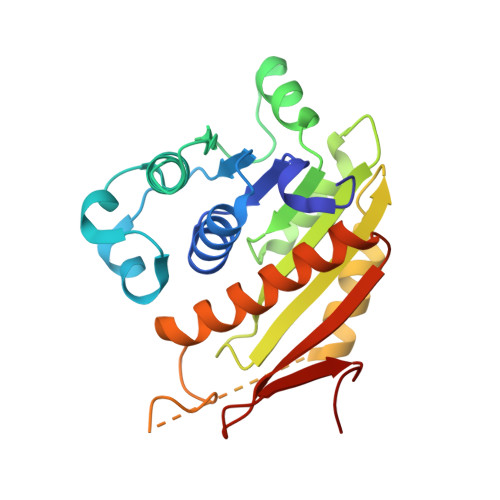
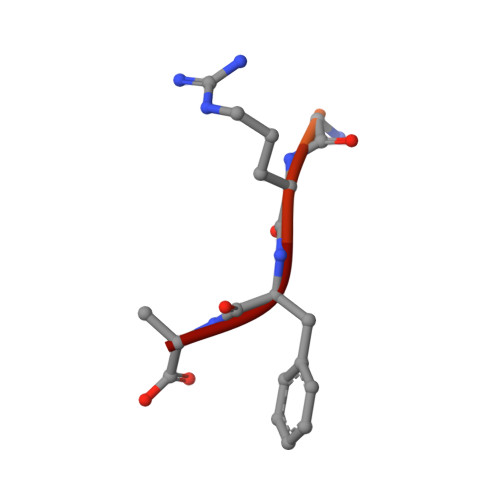
An E1-E2 fusion protein primes antiviral immune signalling in bacteria.
Ledvina, H.E., Ye, Q., Gu, Y., Sullivan, A.E., Quan, Y., Lau, R.K., Zhou, H., Corbett, K.D., Whiteley, A.T.(2023) Nature 616: 319-325
- PubMed: 36755092 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05647-4
- Primary Citation Related Structures:
7TO3, 7TQD, 7TSQ, 7TSX - PubMed Abstract:
In all organisms, innate immune pathways sense infection and rapidly activate potent immune responses while avoiding inappropriate activation (autoimmunity). In humans, the innate immune receptor cyclic GMP-AMP synthase (cGAS) detects viral infection to produce the nucleotide second messenger cyclic GMP-AMP (cGAMP), which initiates stimulator of interferon genes (STING)-dependent antiviral signalling 1 . Bacteria encode evolutionary predecessors of cGAS called cGAS/DncV-like nucleotidyltransferases 2 (CD-NTases), which detect bacteriophage infection and produce diverse nucleotide second messengers 3 . How bacterial CD-NTase activation is controlled remains unknown. Here we show that CD-NTase-associated protein 2 (Cap2) primes bacterial CD-NTases for activation through a ubiquitin transferase-like mechanism. A cryo-electron microscopy structure of the Cap2-CD-NTase complex reveals Cap2 as an all-in-one ubiquitin transferase-like protein, with distinct domains resembling eukaryotic E1 and E2 proteins. The structure captures a reactive-intermediate state with the CD-NTase C terminus positioned in the Cap2 E1 active site and conjugated to AMP. Cap2 conjugates the CD-NTase C terminus to a target molecule that primes the CD-NTase for increased cGAMP production. We further demonstrate that a specific endopeptidase, Cap3, balances Cap2 activity by cleaving CD-NTase-target conjugates. Our data demonstrate that bacteria control immune signalling using an ancient, minimized ubiquitin transferase-like system and provide insight into the evolution of the E1 and E2 machinery across domains of life.
- Department of Biochemistry, University of Colorado Boulder, Boulder, CO, USA.
Organizational Affiliation: