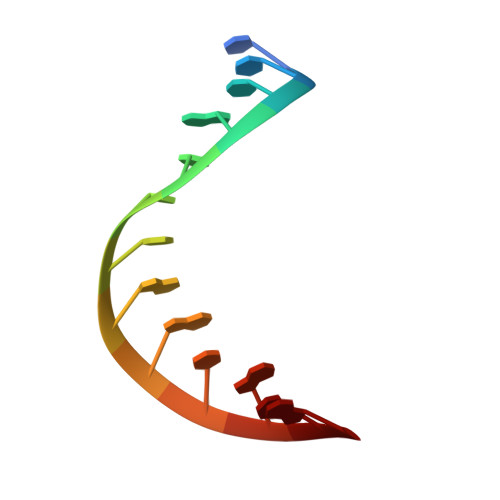
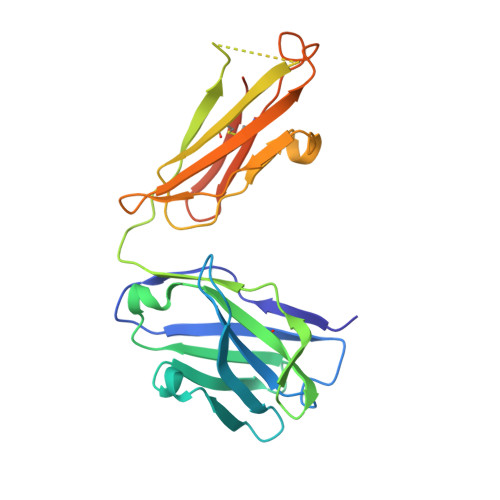
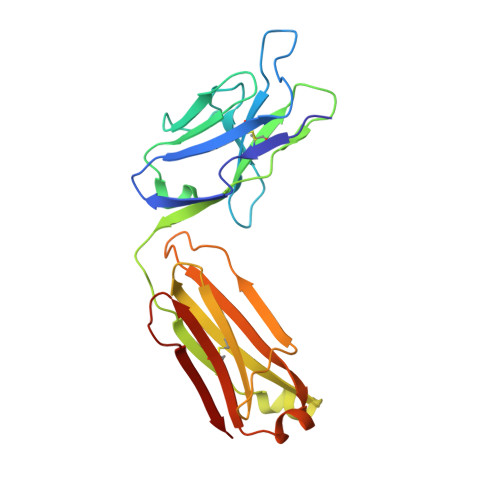
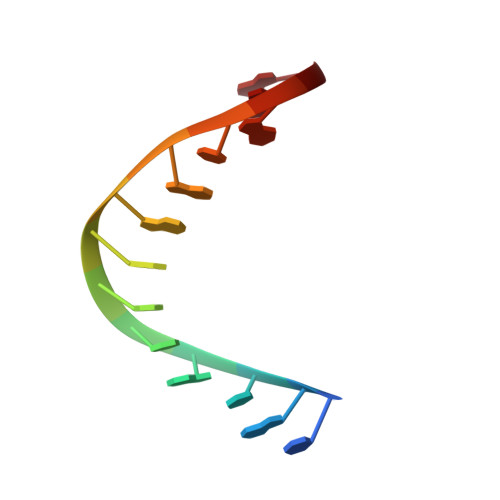
Structural basis of R-loop recognition by the S9.6 monoclonal antibody.
Bou-Nader, C., Bothra, A., Garboczi, D.N., Leppla, S.H., Zhang, J.(2022) Nat Commun 13: 1641-1641
- PubMed: 35347133 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-29187-7
- Primary Citation Related Structures:
7TQA, 7TQB - PubMed Abstract:
R-loops are ubiquitous, dynamic nucleic-acid structures that play fundamental roles in DNA replication and repair, chromatin and transcription regulation, as well as telomere maintenance. The DNA-RNA hybrid-specific S9.6 monoclonal antibody is widely used to map R-loops. Here, we report crystal structures of a S9.6 antigen-binding fragment (Fab) free and bound to a 13-bp hybrid duplex. We demonstrate that S9.6 exhibits robust selectivity in binding hybrids over double-stranded (ds) RNA and in categorically rejecting dsDNA. S9.6 asymmetrically recognizes a compact epitope of two consecutive RNA nucleotides via their 2'-hydroxyl groups and six consecutive DNA nucleotides via their backbone phosphate and deoxyribose groups. Recognition is mediated principally by aromatic and basic residues of the S9.6 heavy chain, which closely track the curvature of the hybrid minor groove. These findings reveal the molecular basis for S9.6 recognition of R-loops, detail its binding specificity, identify a new hybrid-recognition strategy, and provide a framework for S9.6 protein engineering.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, Bethesda, MD, 20892, USA.
Organizational Affiliation: