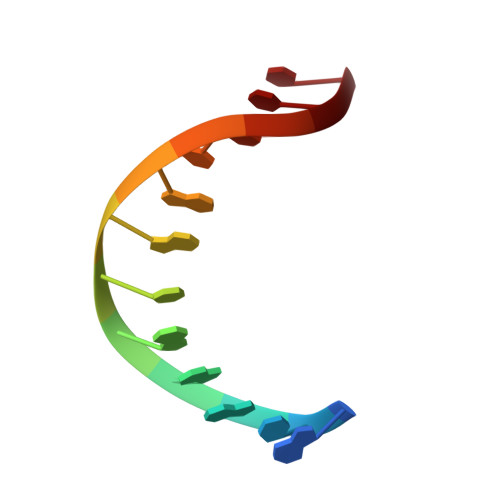
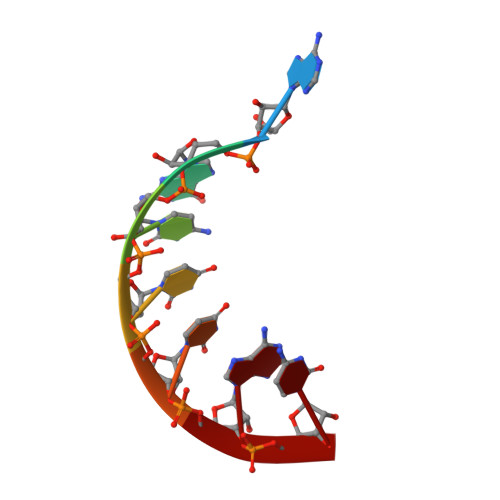
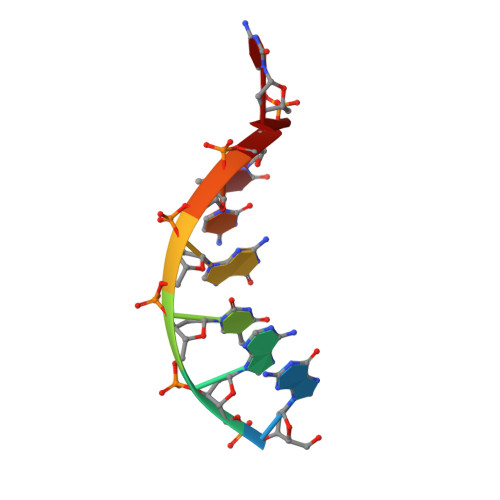
Crystal structure of an RNA/DNA strand exchange junction.
Cofsky, J.C., Knott, G.J., Gee, C.L., Doudna, J.A.(2022) PLoS One 17: e0263547-e0263547
- PubMed: 35436289 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0263547
- Primary Citation Related Structures:
7THB - PubMed Abstract:
Short segments of RNA displace one strand of a DNA duplex during diverse processes including transcription and CRISPR-mediated immunity and genome editing. These strand exchange events involve the intersection of two geometrically distinct helix types-an RNA:DNA hybrid (A-form) and a DNA:DNA homoduplex (B-form). Although previous evidence suggests that these two helices can stack on each other, it is unknown what local geometric adjustments could enable A-on-B stacking. Here we report the X-ray crystal structure of an RNA-5'/DNA-3' strand exchange junction at an anisotropic resolution of 1.6 to 2.2 Å. The structure reveals that the A-to-B helical transition involves a combination of helical axis misalignment, helical axis tilting and compression of the DNA strand within the RNA:DNA helix, where nucleotides exhibit a mixture of A- and B-form geometry. These structural principles explain previous observations of conformational stability in RNA/DNA exchange junctions, enabling a nucleic acid architecture that is repeatedly populated during biological strand exchange events.
- Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, California, United States of America.
Organizational Affiliation: