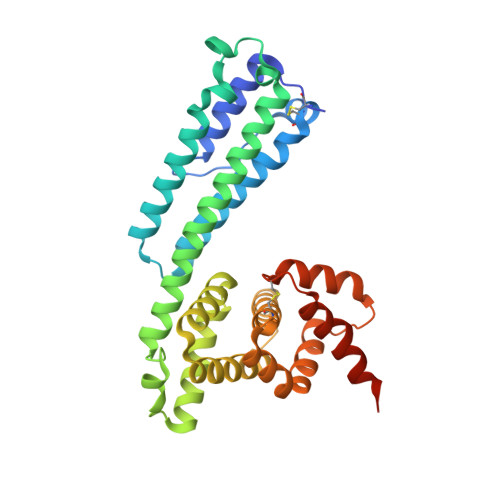
A salivary factor binds a cuticular protein and modulates biting by inducing morphological changes in the mosquito labrum.
Arnoldi, I., Mancini, G., Fumagalli, M., Gastaldi, D., D'Andrea, L., Bandi, C., Di Venere, M., Iadarola, P., Forneris, F., Gabrieli, P.(2022) Curr Biol 32: 3493
- PubMed: 35835123 Search on PubMed
- DOI: https://doi.org/10.1016/j.cub.2022.06.049
- Primary Citation Related Structures:
7TDR, 7TDS - PubMed Abstract:
The mosquito proboscis is an efficient microelectromechanical system, which allows the insect to feed on vertebrate blood quickly and painlessly. Its efficiency is further enhanced by the insect saliva, although through unclear mechanisms. Here, we describe the initial trigger of an unprecedented feedback signaling pathway in Aedes mosquitoes affecting feeding behavior. We identified LIPS proteins in the saliva of Aedes mosquitoes that promote feeding in the vertebrate skin. LIPS show a new all-helical protein fold constituted by two domains. The N-terminal domain interacts with a cuticular protein (Cp19) located at the tip of the mosquito labrum. Upon interaction, the morphology of the labral cuticle changes, and this modification is most likely sensed by proprioceptive neurons. Our study identifies an additional role of mosquito saliva and underlines that the external cuticle is a possible site of key molecular interactions affecting the insect biology and its vector competence.
- The Armenise-Harvard Laboratory of Structural Biology, Department Biology and Biotechnology, University of Pavia, via Ferrata 9, 27100 Pavia, Italy; Entopar lab, Department of Biosciences, University of Milan, via Celoria 26, 20133, Milan, Italy; Centro Interuniversitario di Ricerca sulla Malaria/Italian Malaria Network, Milan, Italy.
Organizational Affiliation: