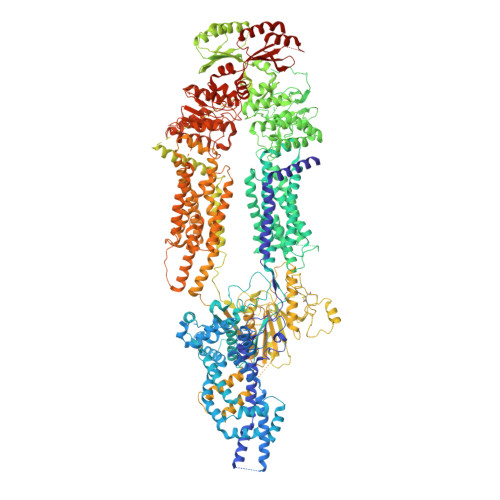
Cholesterol efflux mechanism revealed by structural analysis of human ABCA1 conformational states.
Sun, Y., Li, X.(2022) Nat Cardiovasc Res 1: 238-245
- PubMed: 37181814 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44161-022-00022-y
- Primary Citation Related Structures:
7TBW, 7TBY, 7TBZ, 7TC0 - PubMed Abstract:
ATP-binding cassette transporter A1 (ABCA1) utilizes energy derived from ATP hydrolysis to export cholesterol and phospholipids from macrophages. ABCA1 plays a central role in the biosynthesis of high-density lipoprotein (HDL), which mediates reverse cholesterol transport and prevents detrimental lipid deposition. Mutations in ABCA1 cause Tangier disease characterized by a remarkable reduction in the amount of HDL in blood. Here we present cryo-electron microscopy structures of human ABCA1 in ATP-bound and nucleotide-free states. Structural comparison reveals that ATP molecules pull the nucleotide-binding domains together, inducing movements of transmembrane helices 1, 2, 7 and 8 through a series of salt-bridge interactions. Subsequently, extracellular domains (ECDs) undergo a rotation and introduce conformational changes in the ECD-transmembrane interface. In addition, while we observe a sterol-like molecule in ECDs, no such density was observed in the structure of an HDL-deficiency mutant ABCA1 Y482C , demonstrating the physiological importance of ECDs and a putative interaction mode between ABCA1 and its lipid acceptors. Thus, these structures, along with cholesterol efflux assays, advance the understanding ABCA1-mediated reverse cholesterol transport.
- Department of Molecular Genetics, University of Texas Southwestern Medical Center, Dallas, TX, USA.
Organizational Affiliation: