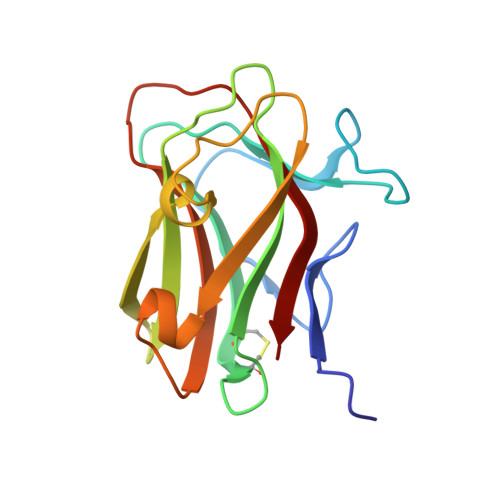
Unique properties of a Dictyostelium discoideum carbohydrate-binding module expand our understanding of CBM-ligand interactions.
Liberato, M.V., Campos, B.M., Tomazetto, G., Crouch, L.I., Garcia, W., Zeri, A.C.M., Bolam, D.N., Squina, F.M.(2022) J Biological Chem 298: 101891-101891
- PubMed: 35378128
- DOI: https://doi.org/10.1016/j.jbc.2022.101891
- Primary Citation Related Structures:
7T7Y, 7T7Z - PubMed Abstract:
Deciphering how enzymes interact, modify, and recognize carbohydrates has long been a topic of interest in academic, pharmaceutical, and industrial research. Carbohydrate-binding modules (CBMs) are noncatalytic globular protein domains attached to carbohydrate-active enzymes that strengthen enzyme affinity to substrates and increase enzymatic efficiency via targeting and proximity effects. CBMs are considered auspicious for various biotechnological purposes in textile, food, and feed industries, representing valuable tools in basic science research and biomedicine. Here, we present the first crystallographic structure of a CBM8 family member (CBM8), DdCBM8, from the slime mold Dictyostelium discoideum, which was identified attached to an endo-β-1,4-glucanase (glycoside hydrolase family 9). We show that the planar carbohydrate-binding site of DdCBM8, composed of aromatic residues, is similar to type A CBMs that are specific for crystalline (multichain) polysaccharides. Accordingly, pull-down assays indicated that DdCBM8 was able to bind insoluble forms of cellulose. However, affinity gel electrophoresis demonstrated that DdCBM8 also bound to soluble (single chain) polysaccharides, especially glucomannan, similar to type B CBMs, although it had no apparent affinity for oligosaccharides. Therefore, the structural characteristics and broad specificity of DdCBM8 represent exceptions to the canonical CBM classification. In addition, mutational analysis identified specific amino acid residues involved in ligand recognition, which are conserved throughout the CBM8 family. This advancement in the structural and functional characterization of CBMs contributes to our understanding of carbohydrate-active enzymes and protein-carbohydrate interactions, pushing forward protein engineering strategies and enhancing the potential biotechnological applications of glycoside hydrolase accessory modules.
- Programa de Processos Tecnológicos e Ambientais, Universidade de Sorocaba (UNISO), Sorocaba, SP, Brazil.
Organizational Affiliation: