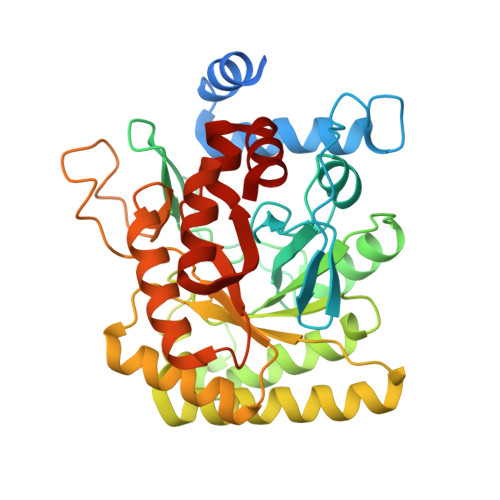
Structural insights into inhibition of the drug target dihydroorotate dehydrogenase by bacterial hydroxyalkylquinolines.
Horwitz, S.M., Blue, T.C., Ambarian, J.A., Hoshino, S., Seyedsayamdost, M.R., Davis, K.M.(2022) RSC Chem Biol 3: 420-425
- PubMed: 35441142 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d1cb00255d
- Primary Citation Related Structures:
7T5K, 7T5Y, 7T6C, 7T6H - PubMed Abstract:
Hydroxyalkylquinolines (HAQs) are ubiquitious natural products but their interactions with associated protein targets remain elusive. We report X-ray crystal structures of two HAQs in complex with dihydroorotate dehydrogenase (DHODH). Our results reveal the structural basis of DHODH inhibition by HAQs and open the door to downstream structure-activity relationship studies.
- Department of Chemistry, Emory University Atlanta GA 30322 USA katherine.davis@emory.edu.
Organizational Affiliation: