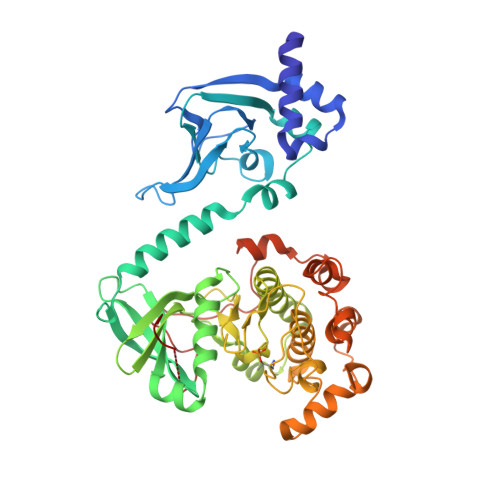
Selective small molecule activation of PKG1alpha: structure and function
Metwally, E., Mak, V., Sylvestre, H.L., McEwan, P., Baker, J.J., Rose, Y., Patel, A., Lim, Y.-H., Healy, D., Hanisak, J., Ermakov, G., Beaumont, M., Cheng, A.C., Greshock, T., Fischmann, T.O.(2023) Commun Biol