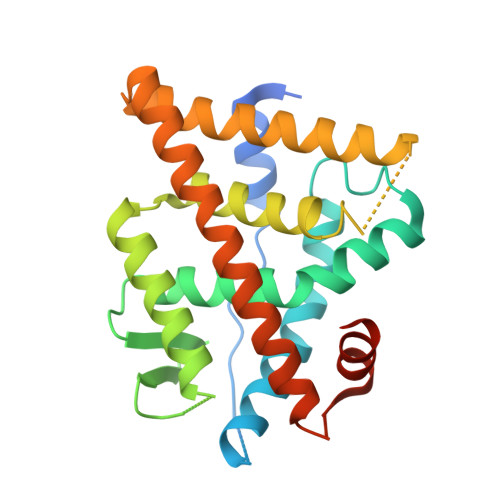
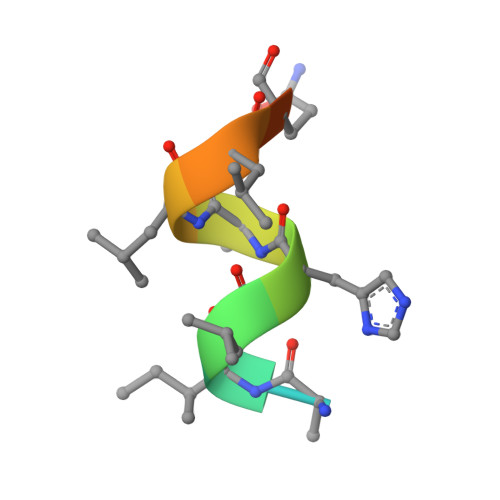A New Chemotype of Chemically Tractable Nonsteroidal Estrogens Based on a Thieno[2,3- d ]pyrimidine Core.
Sammeta, V.R., Norris, J.D., Artham, S., Torrice, C.D., Byemerwa, J., Joiner, C., Fanning, S.W., McDonnell, D.P., Willson, T.M.(2022) ACS Med Chem Lett 13: 1151-1158
- PubMed: 35859859 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.2c00180
- Primary Citation Related Structures:
7RKE, 7T2X - PubMed Abstract:
Despite continued interest in the development of nonsteroidal estrogens and antiestrogens, there are only a few chemotypes of estrogen receptor ligands. Using targeted screening in a ligand sensing assay, we identified a phenolic thieno[2,3- d ]pyrimidine with affinity for estrogen receptor α. An efficient three-step synthesis of the heterocyclic core and structure-guided optimization of the substituents resulted in a series of potent nonsteroidal estrogens. The chemical tractability of the thieno[2,3- d ]pyrimidine chemotype will support the design of new estrogen receptor ligands as therapeutic hormones and antihormones.
- Structural Genomics Consortium and Division of Chemical Biology and Medicinal Chemistry, UNC Eshelman School of Pharmacy, University of North Carolina, Chapel Hill, North Carolina 27599, United States.
Organizational Affiliation: