Sequence of conformation changes of DnaB helicase during DNA unwinding and priming in Escherichia coli
Oakley, A.J., Xu, Z.Q.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Replicative DNA helicase | 471 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: dnaB, groP, grpA, b4052, JW4012 EC: 3.6.4.12 (PDB Primary Data), 5.6.2.3 (UniProt) |  | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0ACB0 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*T)-3') | G [auth M] | 20 | Escherichia coli |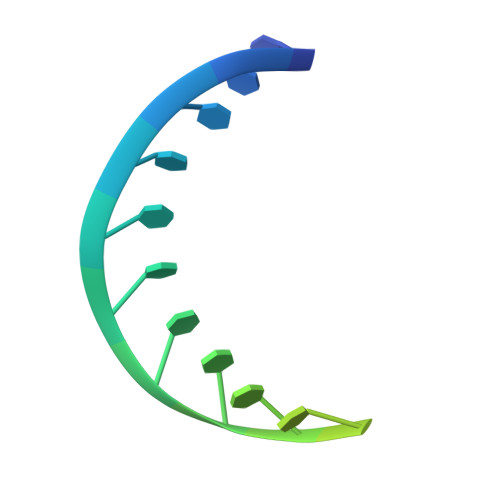  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ANP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | H [auth C], J [auth D], L [auth E], N [auth F] | PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER C10 H17 N6 O12 P3 PVKSNHVPLWYQGJ-KQYNXXCUSA-N |  | ||
| MG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | I [auth C], K [auth D], M [auth E], O [auth F] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.19 |
| RECONSTRUCTION | RELION | 3.07 |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Australian Research Council (ARC) | Australia | DP210100365 |