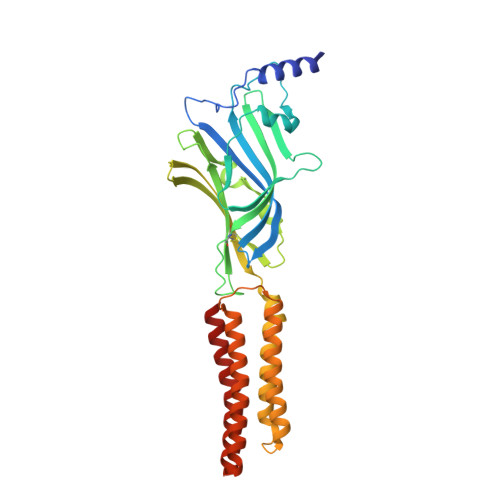
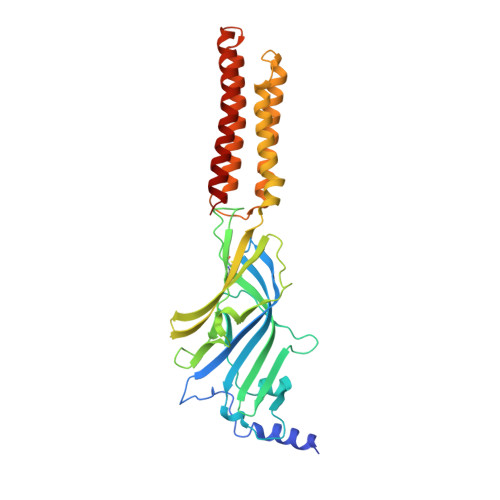
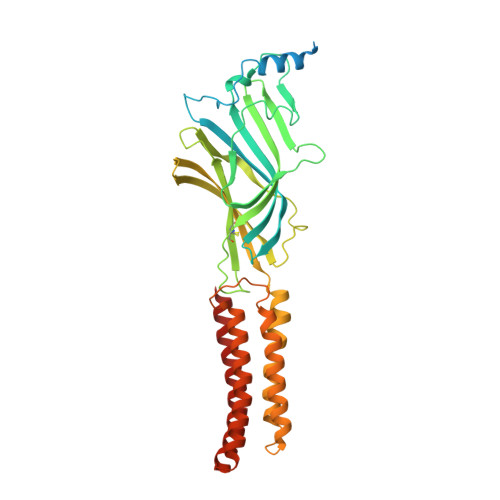
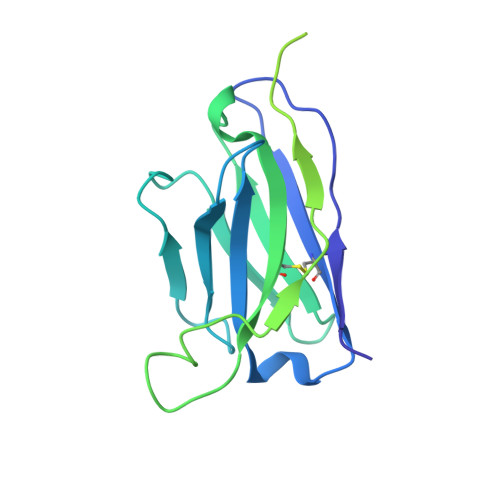
Structural mechanisms of GABA A receptor autoimmune encephalitis.
Noviello, C.M., Kreye, J., Teng, J., Pruss, H., Hibbs, R.E.(2022) Cell 185: 2469
- PubMed: 35803245 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2022.06.025
- Primary Citation Related Structures:
7T0W, 7T0Z - PubMed Abstract:
Autoantibodies targeting neuronal membrane proteins can cause encephalitis, seizures, and severe behavioral abnormalities. While antibodies for several neuronal targets have been identified, structural details on how they regulate function are unknown. Here we determined cryo-electron microscopy structures of antibodies derived from an encephalitis patient bound to the γ-aminobutyric acid type A (GABA A ) receptor. These antibodies induced severe encephalitis by directly inhibiting GABA A function, resulting in nervous-system hyperexcitability. The structures reveal mechanisms of GABA A inhibition and pathology. One antibody directly competes with a neurotransmitter and locks the receptor in a resting-like state. The second antibody targets the subunit interface involved in binding benzodiazepines and antagonizes diazepam potentiation. We identify key residues in these antibodies involved in specificity and affinity and confirm structure-based hypotheses for functional effects using electrophysiology. Together these studies define mechanisms of direct functional antagonism of neurotransmission underlying autoimmune encephalitis in a human patient.
- Department of Neuroscience, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: