Crystal structure of the BCL6 BTB domain in complex with OICR-10562
Kuntz, D.A., Prive, G.G.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Isoform 2 of B-cell lymphoma 6 protein | 127 | Homo sapiens | Mutation(s): 3 Gene Names: BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | 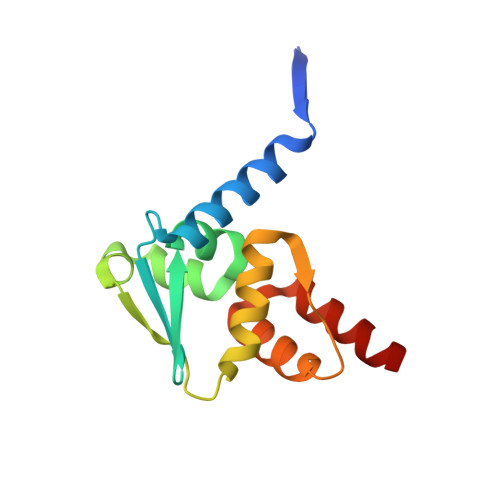 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P41182 GTEx: ENSG00000113916 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P41182 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| E1I (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | H [auth A] I [auth B] J [auth C] K [auth D] L [auth E] | N-[5-chloro-2-(morpholin-4-yl)pyridin-4-yl]-2-[5-(3-cyano-4-hydroxy-5-methylphenyl)-3-methyl-2-(1-methyl-1H-pyrazol-4-yl)-4-oxo-3,4-dihydro-7H-pyrrolo[2,3-d]pyrimidin-7-yl]acetamide C30 H28 Cl N9 O4 DZAOOXDSLZEBAB-UHFFFAOYSA-N |  | ||
| DMS Download:Ideal Coordinates CCD File | G [auth A], M [auth E] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 62.592 | α = 90 |
| b = 102.549 | β = 115.48 |
| c = 65.028 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ontario Institute for Cancer Research | Canada | -- |