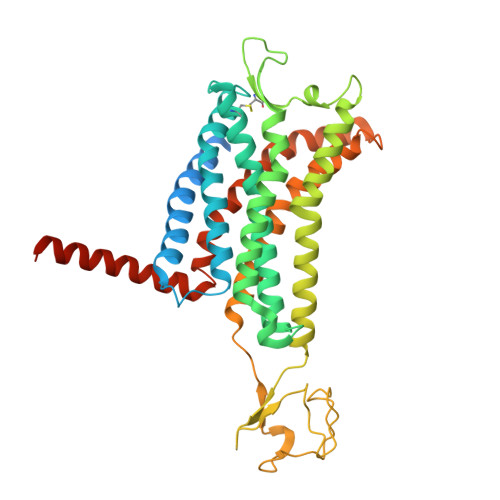
Structural insight into apelin receptor-G protein stoichiometry.
Yue, Y., Liu, L., Wu, L.J., Wu, Y., Wang, L., Li, F., Liu, J., Han, G.W., Chen, B., Lin, X., Brouillette, R.L., Breault, E., Longpre, J.M., Shi, S., Lei, H., Sarret, P., Stevens, R.C., Hanson, M.A., Xu, F.(2022) Nat Struct Mol Biol 29: 688-697
- PubMed: 35817871 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00797-5
- Primary Citation Related Structures:
7SUS, 7W0L, 7W0M, 7W0N, 7W0O, 7W0P - PubMed Abstract:
The technique of cryogenic-electron microscopy (cryo-EM) has revolutionized the field of membrane protein structure and function with a focus on the dominantly observed molecular species. This report describes the structural characterization of a fully active human apelin receptor (APJR) complexed with heterotrimeric G protein observed in both 2:1 and 1:1 stoichiometric ratios. We use cryo-EM single-particle analysis to determine the structural details of both species from the same sample preparation. Protein preparations, in the presence of the endogenous peptide ligand ELA or a synthetic small molecule, both demonstrate these mixed stoichiometric states. Structural differences in G protein engagement between dimeric and monomeric APJR suggest a role for the stoichiometry of G protein-coupled receptor- (GPCR-)G protein coupling on downstream signaling and receptor pharmacology. Furthermore, a small, hydrophobic dimer interface provides a starting framework for additional class A GPCR dimerization studies. Together, these findings uncover a mechanism of versatile regulation through oligomerization by which GPCRs can modulate their signaling.
- iHuman Institute, ShanghaiTech University, Shanghai, China.
Organizational Affiliation: