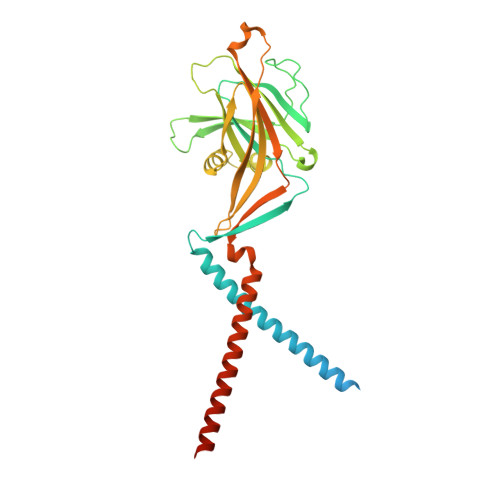
Gating choreography and mechanism of the human proton-activated chloride channel ASOR.
Wang, C., Polovitskaya, M.M., Delgado, B.D., Jentsch, T.J., Long, S.B.(2022) Sci Adv 8: eabm3942-eabm3942
- PubMed: 35108041 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abm3942
- Primary Citation Related Structures:
7SQF, 7SQG, 7SQH - PubMed Abstract:
The proton-activated chloride channel ASOR (TMEM206/PAC) permeates anions across cellular membranes in response to acidification, thereby enhancing acid-induced cell death and regulating endocytosis. The molecular mechanisms of pH-dependent control are not understood, in part because structural information for an activated conformation of ASOR is lacking. Here, we reconstitute function from purified protein and present a 3.1-Å-resolution cryo-electron microscopy structure of human ASOR at acidic pH in an activated conformation. The work contextualizes a previous acidic pH structure as a desensitized conformation. Combined with electrophysiological studies and high-resolution structures of resting and desensitized states, the work reveals mechanisms of proton sensing and ion pore gating. Clusters of extracellular acidic residues function as pH sensors and coalesce when protonated. Ensuing conformational changes induce metamorphosis of transmembrane helices to fashion an ion conduction pathway unique to the activated conformation. The studies identify a new paradigm of channel gating in this ubiquitous ion channel.
- Structural Biology Program, Memorial Sloan Kettering Cancer Center, 1275 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: