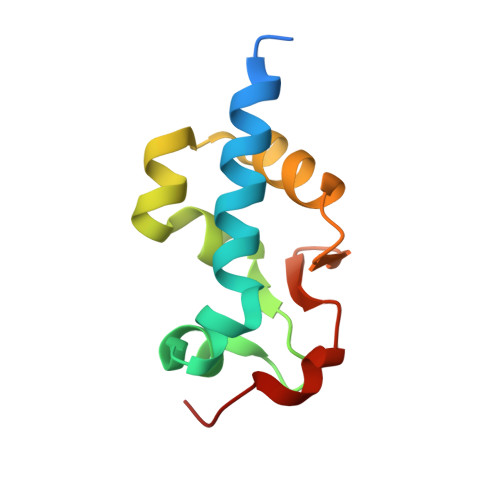
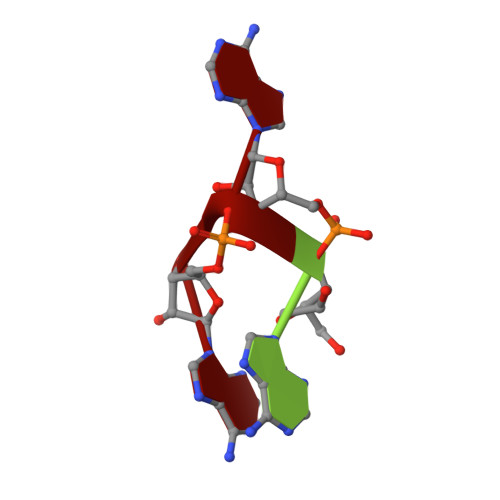
Structural basis of 3'-end poly(A) RNA recognition by LARP1.
Kozlov, G., Mattijssen, S., Jiang, J., Nyandwi, S., Sprules, T., Iben, J.R., Coon, S.L., Gaidamakov, S., Noronha, A.M., Wilds, C.J., Maraia, R.J., Gehring, K.(2022) Nucleic Acids Res 50: 9534-9547
- PubMed: 35979957 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac696
- Primary Citation Related Structures:
7SOO, 7SOP, 7SOQ, 7SOR, 7SOS, 7SOT, 7SOU, 7SOV - PubMed Abstract:
La-related proteins (LARPs) comprise a family of RNA-binding proteins involved in a wide range of posttranscriptional regulatory activities. LARPs share a unique tandem of two RNA-binding domains, La motif (LaM) and RNA recognition motif (RRM), together referred to as a La-module, but vary in member-specific regions. Prior structural studies of La-modules reveal they are pliable platforms for RNA recognition in diverse contexts. Here, we characterize the La-module of LARP1, which plays an important role in regulating synthesis of ribosomal proteins in response to mTOR signaling and mRNA stabilization. LARP1 has been well characterized functionally but no structural information exists for its La-module. We show that unlike other LARPs, the La-module in LARP1 does not contain an RRM domain. The LaM alone is sufficient for binding poly(A) RNA with submicromolar affinity and specificity. Multiple high-resolution crystal structures of the LARP1 LaM domain in complex with poly(A) show that it is highly specific for the RNA 3'-end, and identify LaM residues Q333, Y336 and F348 as the most critical for binding. Use of a quantitative mRNA stabilization assay and poly(A) tail-sequencing demonstrate functional relevance of LARP1 RNA binding in cells and provide novel insight into its poly(A) 3' protection activity.
- Department of Biochemistry, McGill University, Montréal, Canada.
Organizational Affiliation: