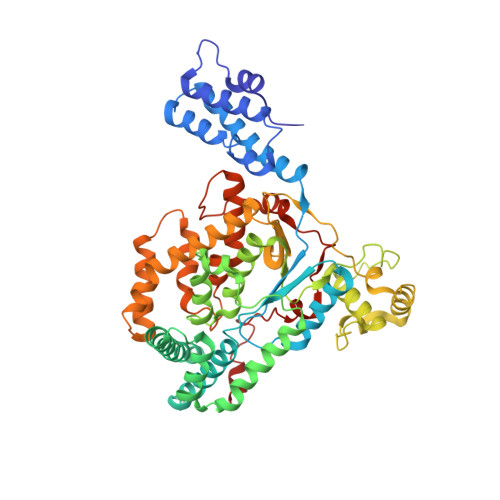
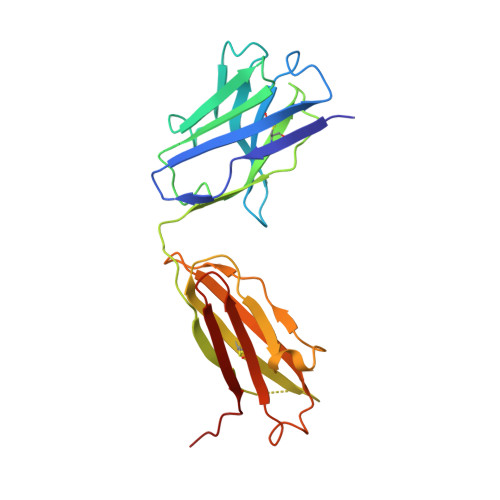
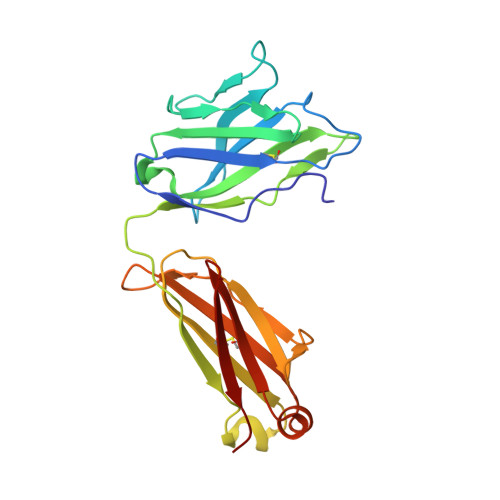
Novel structural insights for a pair of monoclonal antibodies recognizing non-overlapping epitopes of the glucosyltransferase domain of Clostridium difficile toxin B.
Liu, J., Kothe, M., Zhang, J., Oloo, E., Stegalkina, S., Mundle, S.T., Li, L., Zhang, J., Cole, L.E., Barone, L., Biemann, H.P., Kleanthous, H., Anosova, N.G., Anderson, S.F.(2022) Curr Res Struct Biol 4: 96-105
- PubMed: 35469152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.crstbi.2022.03.003
- Primary Citation Related Structures:
7SO5, 7SO7 - PubMed Abstract:
Clostridium difficile toxins are the primary causative agents for hospital-acquired diarrhea and pseudomembranous colitis. Numerous monoclonal antibodies (mAbs) targeting different domains of Clostridium difficile toxin have been reported. Here we report the crystal structures of two mAbs, B1 and B2, in complex with the glycosyltransferase domain (GTD) of the Clostridium difficile toxin B (TcdB). B2 bound to the N-terminal 4 helix bundle of the GTD, a conserved membrane localization domain (MLD) found in the large clostridial glycosylating toxin family implicated in targeting plasma membrane. B1 bound to a distinct epitope at the hinge region between the MLD and the catalytic subdomain of the GTD. Functional studies revealed the potency of these mAbs in vitro and in vivo to be synergistic when given in combination.
- Sanofi Integrated Drug Discovery, 153 2nd Ave, Waltham, 02451, USA.
Organizational Affiliation: