Crystal Structure of Siderophore Binding Protein FatB from Desulfitobacterium hafniense
Kim, Y., Patel, H.P., Nordquist, K.A., Schaab, K.M., Sha, J., Babnigg, G., Bond, A.H., Joachimiak, A., Midwest Center for Structural GenomicsTo be published.
Experimental Data Snapshot
Starting Model: in silico
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ferric-anguibactin-binding protein FatB | 314 | Desulfitobacterium hafniense DCB-2 | Mutation(s): 0 Gene Names: Dhaf_2916 | 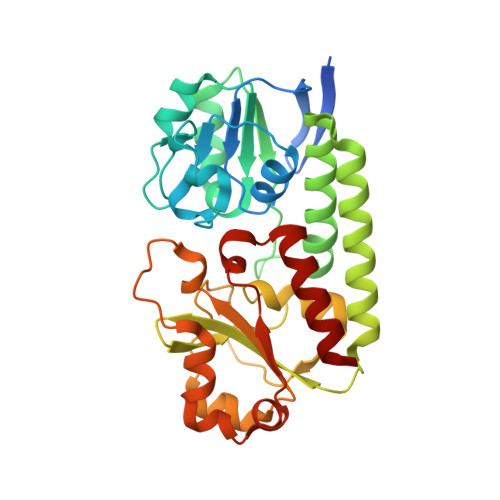 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q24WN9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| DBS (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | B [auth A], C [auth A] | 2-(2,3-DIHYDROXY-BENZOYLAMINO)-3-HYDROXY-PROPIONIC ACID C10 H11 N O6 VDTYHTVHFIIEIL-LURJTMIESA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | D [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | E [auth A], F [auth A], G [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| FE Download:Ideal Coordinates CCD File | H [auth A] | FE (III) ION Fe VTLYFUHAOXGGBS-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | I [auth A], J [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 78.884 | α = 90 |
| b = 78.884 | β = 90 |
| c = 122.402 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| HKL-3000 | phasing |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | -- |