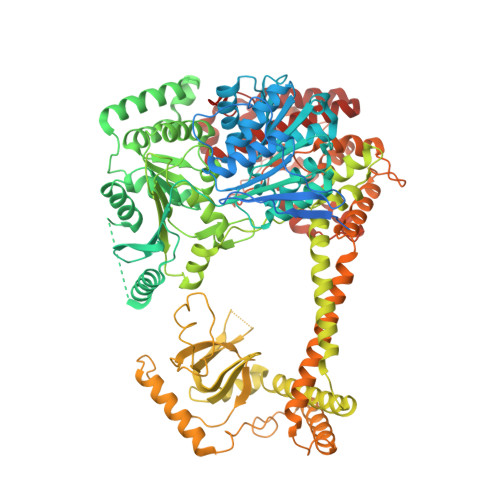
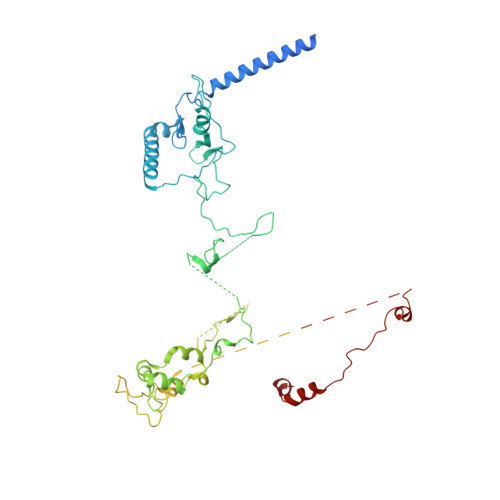
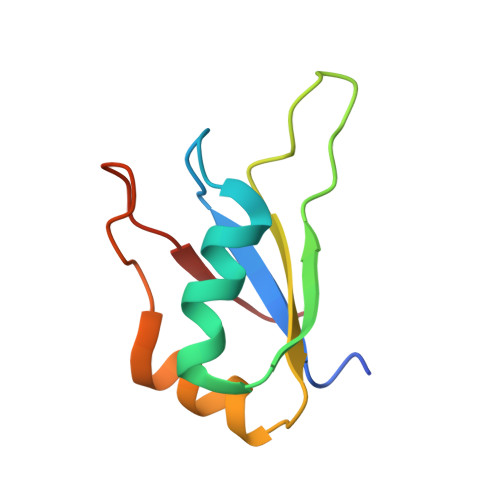
Structural basis for RNA surveillance by the human nuclear exosome targeting (NEXT) complex.
Puno, M.R., Lima, C.D.(2022) Cell 185: 2132-2147.e26
- PubMed: 35688134
- DOI: https://doi.org/10.1016/j.cell.2022.04.016
- Primary Citation of Related Structures:
7S7B, 7S7C - PubMed Abstract:
RNA quality control relies on co-factors and adaptors to identify and prepare substrates for degradation by ribonucleases such as the 3' to 5' ribonucleolytic RNA exosome. Here, we determined cryogenic electron microscopy structures of human nuclear exosome targeting (NEXT) complexes bound to RNA that reveal mechanistic insights to substrate recognition and early steps that precede RNA handover to the exosome. The structures illuminate ZCCHC8 as a scaffold, mediating homodimerization while embracing the MTR4 helicase and flexibly anchoring RBM7 to the helicase core. All three subunits collaborate to bind the RNA, with RBM7 and ZCCHC8 surveying sequences upstream of the 3' end to facilitate RNA capture by MTR4. ZCCHC8 obscures MTR4 surfaces important for RNA binding and extrusion as well as MPP6-dependent recruitment and docking onto the RNA exosome core, interactions that contribute to RNA surveillance by coordinating RNA capture, translocation, and extrusion from the helicase to the exosome for decay.
- Structural Biology Program, Sloan Kettering Institute, Memorial Sloan Kettering Cancer Center, 1275 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: