Structure of a Cell Entry Defective Human Adenovirus Provides Insights into Precursor Proteins and Capsid Maturation.
Yu, X., Mullen, T.M., Abrishami, V., Huiskonen, J.T., Nemerow, G.R., Reddy, V.S.(2021) J Mol Biology 434: 167350-167350
- PubMed: 34774568 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2021.167350
- Primary Citation Related Structures:
7S78 - PubMed Abstract:
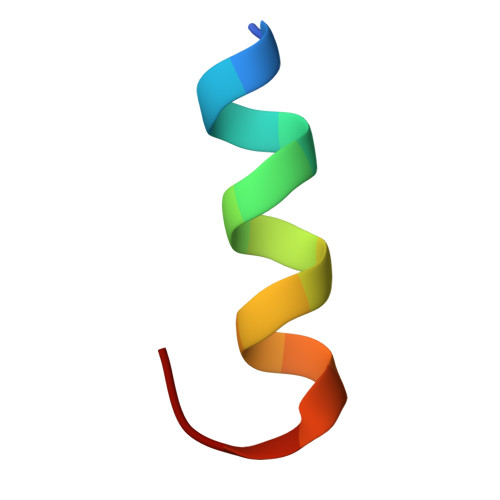
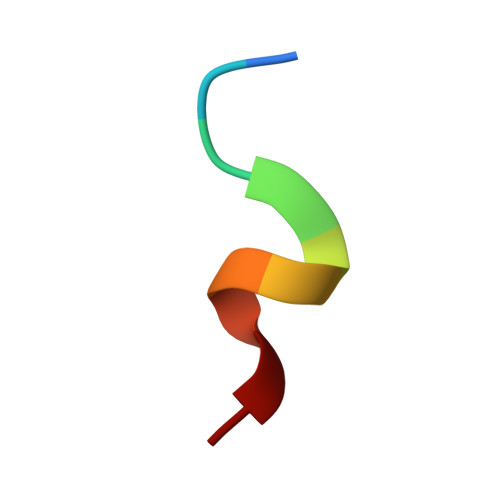
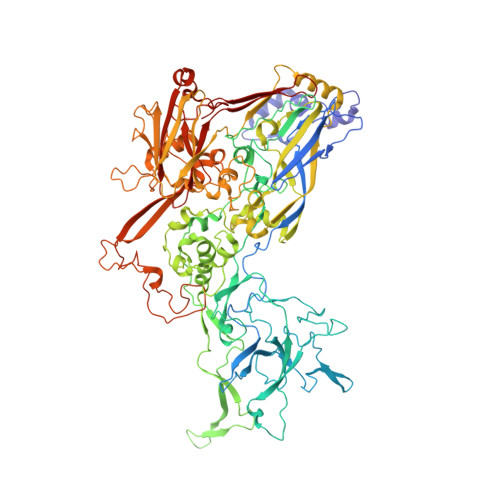
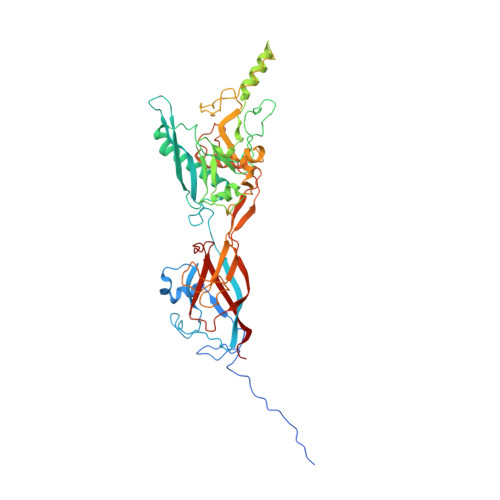
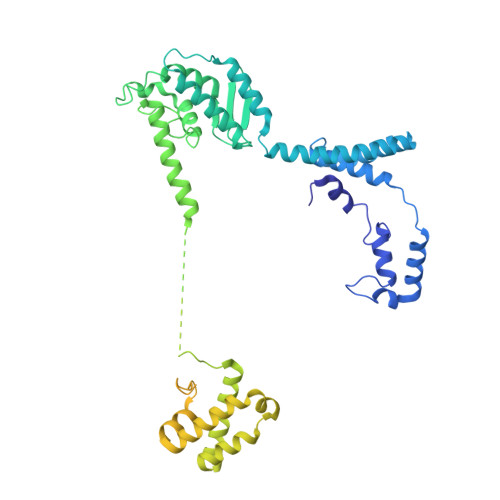
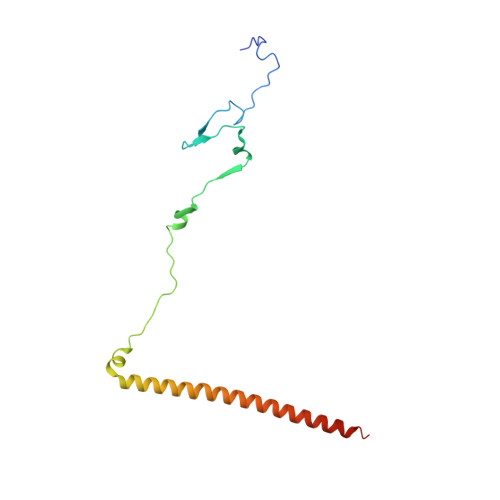
Maturation of adenoviruses is distinguished by proteolytic processing of several interior minor capsid proteins and core proteins by the adenoviral protease and subsequent reorganization of adenovirus core. We report the results derived from the icosahedrally averaged cryo-EM structure of a cell entry defective form of adenovirus, designated ts1, at a resolution of 3.7 Å as well as of the localized reconstructions of unique hexons and penton base. The virion structure revealed the structures and organization of precursors of minor capsid proteins, pIIIa, pVI and pVIII, which are closely associated with the hexons on the capsid interior. In addition to a well-ordered helical domain (a.a. 310-397) of pIIIa, highlights of the structure include the precursors of VIII display significantly different structures near the cleavage sites. Moreover, we traced residues 4-96 of the membrane lytic protein (pVI) that includes an amphipathic helix occluded deep in the hexon cavity suggesting the possibility of co-assembly of hexons with the precursors of VI. In addition, we observe a second copy of pVI ordered up to residue L40 in the peripentonal hexons and a few fragments of density corresponding to 2nd and 3rd copies of pVI in other hexons. However, we see no evidence of precursors of VII binding in the hexon cavity. These findings suggest the possibility that differently bound pVI molecules undergo processing at the N-terminal cleavage sites at varying efficiencies, subsequently creating competition between the cleaved and uncleaved forms of VI, followed by reorganization, processing, and release of VI molecules from the hexon cavities.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: