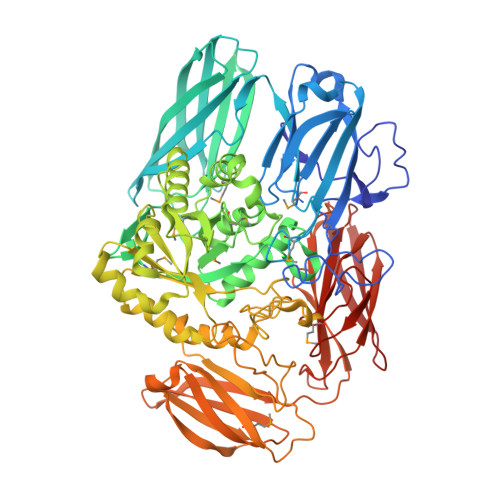
The crystal structure from microfluidic crystals of glycosyl hydrolase family 2 (GH2) member from Bacteroides cellulosilyticus
Kim, Y., Nocek, B., Endres, M., Joachimiak, G., Johnson, J., Babnigg, G., Joachimiak, A., Midwest Center for Structural Genomics (MCSG)To be published.