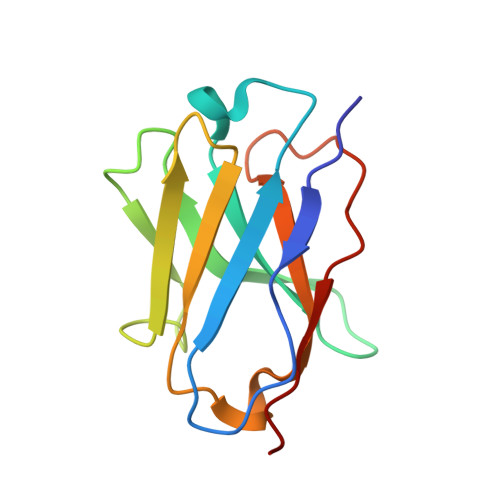
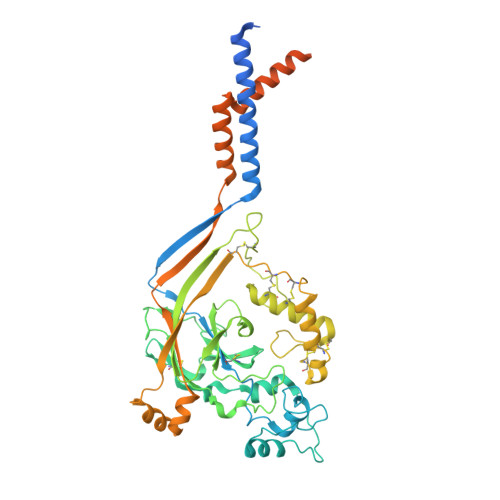
Structure and analysis of nanobody binding to the human ASIC1a ion channel.
Wu, Y., Chen, Z., Sigworth, F.J., Canessa, C.M.(2021) Elife 10
- PubMed: 34319232 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.67115
- Primary Citation Related Structures:
7RNN - PubMed Abstract:
ASIC1a is a proton-gated sodium channel involved in modulation of pain, fear, addiction, and ischemia-induced neuronal injury. We report isolation and characterization of alpaca-derived nanobodies (Nbs) that specifically target human ASIC1a. Cryo-electron microscopy of the human ASIC1a channel at pH 7.4 in complex with one of these, Nb.C1, yielded a structure at 2.9 Å resolution. It is revealed that Nb.C1 binds to a site overlapping with that of the Texas coral snake toxin (MitTx1) and the black mamba venom Mambalgin-1; however, the Nb.C1-binding site does not overlap with that of the inhibitory tarantula toxin psalmotoxin-1 (PcTx1). Fusion of Nb.C1 with PcTx1 in a single polypeptide markedly enhances the potency of PcTx1, whereas competition of Nb.C1 and MitTx1 for binding reduces channel activation by the toxin. Thus, Nb.C1 is a molecular tool for biochemical and structural studies of hASIC1a; a potential antidote to the pain-inducing component of coral snake bite; and a candidate to potentiate PcTx1-mediated inhibition of hASIC1a in vivo for therapeutic applications.
- Basic Sciences Department, Tsinghua University School of Medicine, Beijing, China.
Organizational Affiliation: