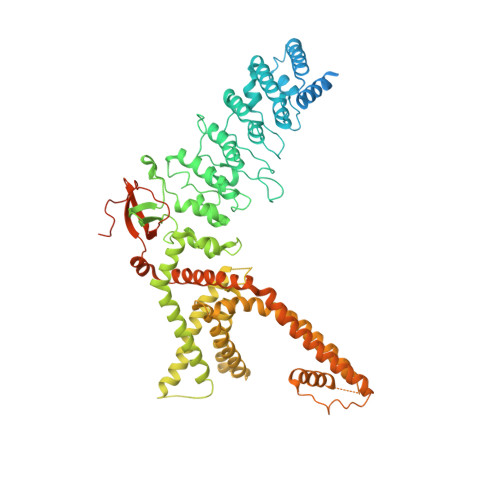
Structural mechanism of TRPV3 channel inhibition by the plant-derived coumarin osthole.
Neuberger, A., Nadezhdin, K.D., Zakharian, E., Sobolevsky, A.I.(2021) EMBO Rep 22: e53233-e53233
- PubMed: 34472684 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embr.202153233
- Primary Citation Related Structures:
7RAS, 7RAU - PubMed Abstract:
TRPV3, a representative of the vanilloid subfamily of TRP channels, is predominantly expressed in skin keratinocytes and has been implicated in cutaneous sensation and associated with numerous skin pathologies and cancers. TRPV3 is inhibited by the natural coumarin derivative osthole, an active ingredient of Cnidium monnieri, which has been used in traditional Chinese medicine for the treatment of a variety of human diseases. However, the structural basis of channel inhibition by osthole has remained elusive. Here we present cryo-EM structures of TRPV3 in complex with osthole, revealing two types of osthole binding sites in the transmembrane region of TRPV3 that coincide with the binding sites of agonist 2-APB. Osthole binding converts the channel pore into a previously unidentified conformation with a widely open selectivity filter and closed intracellular gate. Our structures provide insight into competitive inhibition of TRPV3 by osthole and can serve as a template for the design of osthole chemistry-inspired drugs targeting TRPV3-associated diseases.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY, USA.
Organizational Affiliation: