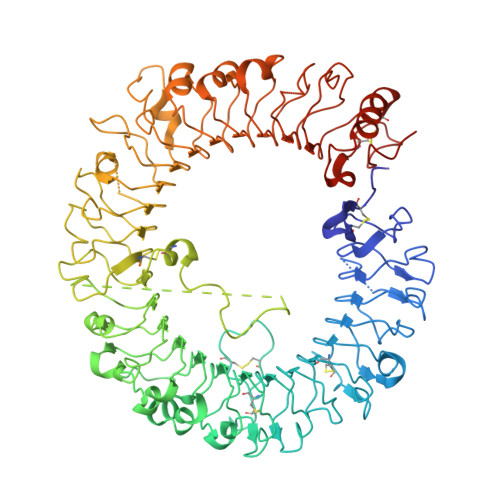
Structure-Based Optimization of a Fragment-like TLR8 Binding Screening Hit to an In Vivo Efficacious TLR7/8 Antagonist.
Betschart, C., Faller, M., Zink, F., Hemmig, R., Blank, J., Vangrevelinghe, E., Bourrel, M., Glatthar, R., Behnke, D., Barker, K., Heizmann, A., Angst, D., Nimsgern, P., Jacquier, S., Junt, T., Zipfel, G., Ruzzante, G., Loetscher, P., Limonta, S., Hawtin, S., Andre, C.B., Boulay, T., Feifel, R., Knoepfel, T.(2022) ACS Med Chem Lett 13: 658-664
- PubMed: 35450354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.1c00696
- Primary Citation Related Structures:
7R52, 7R53, 7R54 - PubMed Abstract:
Inappropriate activation of TLR7 and TLR8 is linked to several autoimmune diseases, such as lupus erythematosus. Here we report on the efficient structure-based optimization of the inhibition of TLR8, starting from a co-crystal structure of a small screening hit. Further optimization of the physicochemical properties for cellular potency and expansion of the structure-activity relationship for dual potency finally resulted in a highly potent TLR7/8 antagonist with demonstrated in vivo efficacy after oral dosing.
- Global Discovery Chemistry, Novartis Institutes for BioMedical Research, CH-4002 Basel, Switzerland.
Organizational Affiliation: