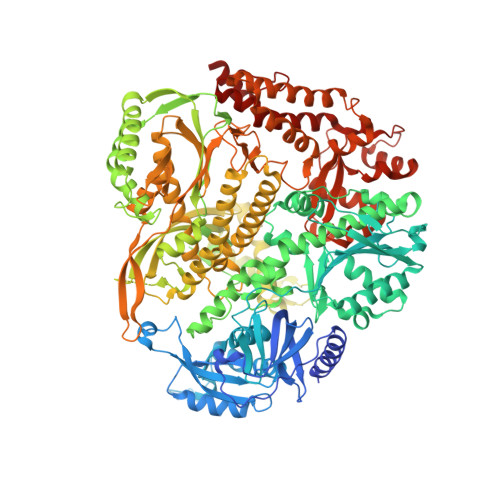
Enhanced polymerase activity permits efficient synthesis by cancer-associated DNA polymerase ε variants at low dNTP levels.
Barbari, S.R., Beach, A.K., Markgren, J.G., Parkash, V., Moore, E.A., Johansson, E., Shcherbakova, P.V.(2022) Nucleic Acids Res 50: 8023-8040
- PubMed: 35822874 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac602
- Primary Citation Related Structures:
7R3X, 7R3Y - PubMed Abstract:
Amino acid substitutions in the exonuclease domain of DNA polymerase ϵ (Polϵ) cause ultramutated tumors. Studies in model organisms suggested pathogenic mechanisms distinct from a simple loss of exonuclease. These mechanisms remain unclear for most recurrent Polϵ mutations. Particularly, the highly prevalent V411L variant remained a long-standing puzzle with no detectable mutator effect in yeast despite the unequivocal association with ultramutation in cancers. Using purified four-subunit yeast Polϵ, we assessed the consequences of substitutions mimicking human V411L, S459F, F367S, L424V and D275V. While the effects on exonuclease activity vary widely, all common cancer-associated variants have increased DNA polymerase activity. Notably, the analog of Polϵ-V411L is among the strongest polymerases, and structural analysis suggests defective polymerase-to-exonuclease site switching. We further show that the V411L analog produces a robust mutator phenotype in strains that lack mismatch repair, indicating a high rate of replication errors. Lastly, unlike wild-type and exonuclease-dead Polϵ, hyperactive variants efficiently synthesize DNA at low dNTP concentrations. We propose that this characteristic could promote cancer cell survival and preferential participation of mutator polymerases in replication during metabolic stress. Our results support the notion that polymerase fitness, rather than low fidelity alone, is an important determinant of variant pathogenicity.
- Eppley Institute for Research in Cancer and Allied Diseases, Fred & Pamela Buffett Cancer Center, University of Nebraska Medical Center, Omaha, NE 68198, USA.
Organizational Affiliation: