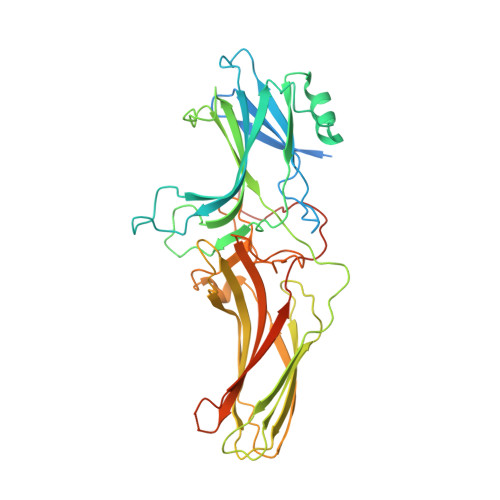
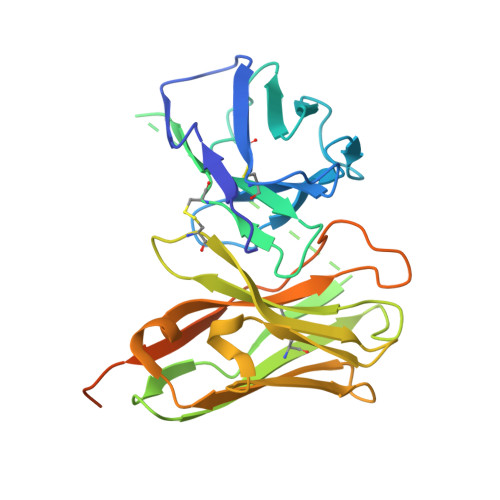
Structure of the vasopressin hormone-V2 receptor-beta-arrestin1 ternary complex.
Bous, J., Fouillen, A., Orcel, H., Trapani, S., Cong, X., Fontanel, S., Saint-Paul, J., Lai-Kee-Him, J., Urbach, S., Sibille, N., Sounier, R., Granier, S., Mouillac, B., Bron, P.(2022) Sci Adv 8: eabo7761-eabo7761
- PubMed: 36054364 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abo7761
- Primary Citation Related Structures:
7R0C, 7R0J - PubMed Abstract:
Arrestins interact with G protein-coupled receptors (GPCRs) to stop G protein activation and to initiate key signaling pathways. Recent structural studies shed light on the molecular mechanisms involved in GPCR-arrestin coupling, but whether this process is conserved among GPCRs is poorly understood. Here, we report the cryo-electron microscopy active structure of the wild-type arginine-vasopressin V2 receptor (V2R) in complex with β-arrestin1. It reveals an atypical position of β-arrestin1 compared to previously described GPCR-arrestin assemblies, associated with an original V2R/β-arrestin1 interface involving all receptor intracellular loops. Phosphorylated sites of the V2R carboxyl terminus are clearly identified and interact extensively with the β-arrestin1 N-lobe, in agreement with structural data obtained with chimeric or synthetic systems. Overall, these findings highlight a notable structural variability among GPCR-arrestin signaling complexes.
- CBS (Centre de Biologie Structurale), Université de Montpellier, CNRS, INSERM, Montpellier, France.
Organizational Affiliation: