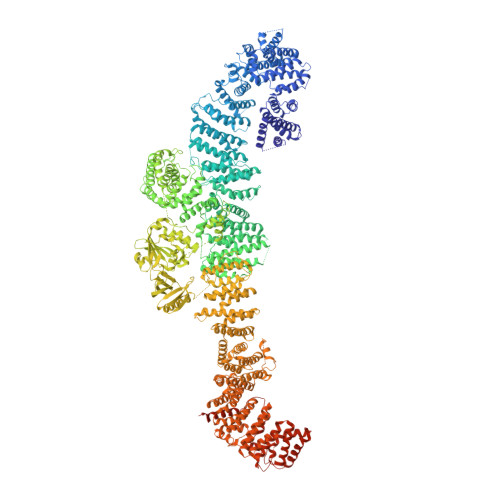
Structural basis of activation of the tumor suppressor protein neurofibromin.
Chaker-Margot, M., Werten, S., Dunzendorfer-Matt, T., Lechner, S., Ruepp, A., Scheffzek, K., Maier, T.(2022) Mol Cell 82: 1288
- PubMed: 35353986 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2022.03.011
- Primary Citation Related Structures:
7R03, 7R04 - PubMed Abstract:
Mutations in the NF1 gene cause the familial genetic disease neurofibromatosis type I, as well as predisposition to cancer. The NF1 gene product, neurofibromin, is a GTPase-activating protein and acts as a tumor suppressor by negatively regulating the small GTPase, Ras. However, structural insights into neurofibromin activation remain incompletely defined. Here, we provide cryoelectron microscopy (cryo-EM) structures that reveal an extended neurofibromin homodimer in two functional states: an auto-inhibited state with occluded Ras-binding site and an asymmetric open state with an exposed Ras-binding site. Mechanistically, the transition to the active conformation is stimulated by nucleotide binding, which releases a lock that tethers the catalytic domain to an extended helical repeat scaffold in the occluded state. Structure-guided mutational analysis supports functional relevance of allosteric control. Disease-causing mutations are mapped and primarily impact neurofibromin stability. Our findings suggest a role for nucleotides in neurofibromin regulation and may lead to therapeutic modulation of Ras signaling.
- Biozentrum, University of Basel, 4056 Basel, Switzerland.
Organizational Affiliation: