Structure of M.BseCI DNA methyltransferase from Geobacillus stearothermophilus.
Mitsikas, D.A., Kouyianou, K., Kotsifaki, D., Providaki, M., Bouriotis, V., Glykos, N.M., Kokkinidis, M.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Modification methylase BseCI | 585 | Geobacillus stearothermophilus | Mutation(s): 0 Gene Names: bseCIM EC: 2.1.1.72 | 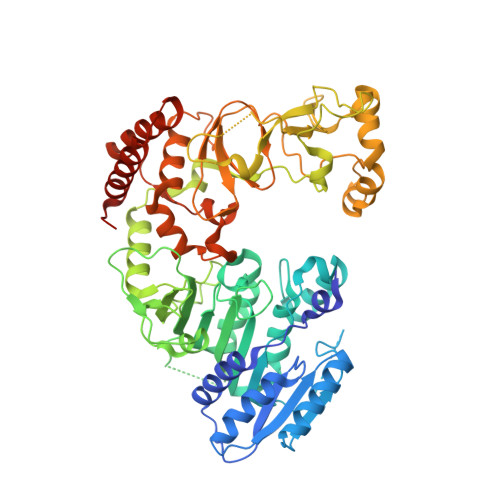 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P43423 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| Unmethylated DNA duplex | B [auth Z] | 10 | Geobacillus stearothermophilus | 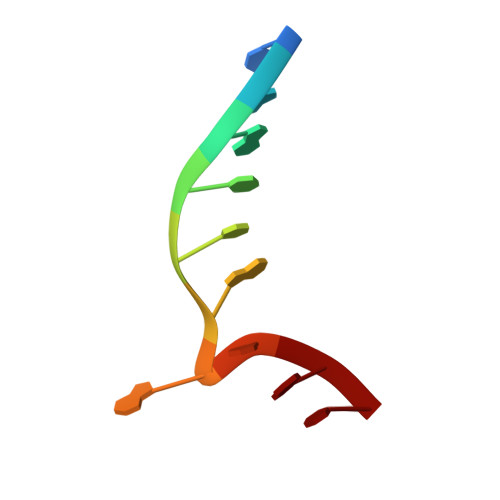 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| Unmethylated DNA duplex | C [auth Y] | 10 | Geobacillus stearothermophilus | 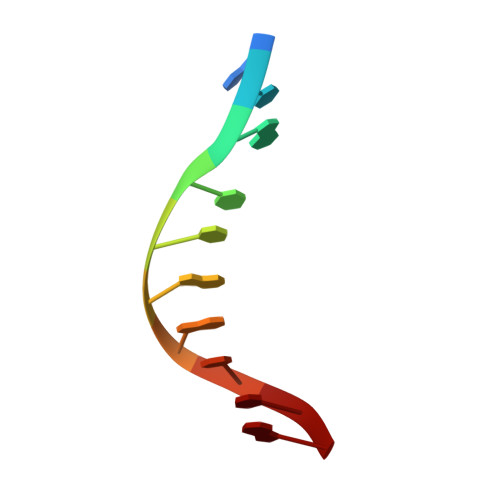 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SAH (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | S-ADENOSYL-L-HOMOCYSTEINE C14 H20 N6 O5 S ZJUKTBDSGOFHSH-WFMPWKQPSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 87.04 | α = 90 |
| b = 87.04 | β = 90 |
| c = 156.35 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| iMOSFLM | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| CRANK | phasing |
| BUCCANEER | model building |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | Greece | -- |