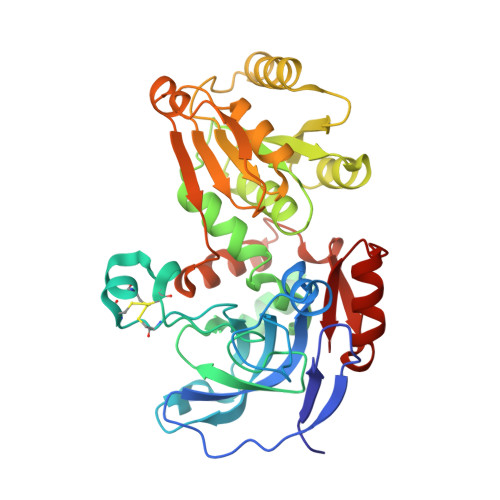
Advanced Insights into Catalytic and Structural Features of the Zinc-Dependent Alcohol Dehydrogenase from Thauera aromatica.
Stark, F., Loderer, C., Petchey, M., Grogan, G., Ansorge-Schumacher, M.B.(2022) Chembiochem 23: e202200149-e202200149
- PubMed: 35557486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cbic.202200149
- Primary Citation Related Structures:
7QUL, 7QUY - PubMed Abstract:
The asymmetric reduction of ketones to chiral hydroxyl compounds by alcohol dehydrogenases (ADHs) is an established strategy for the provision of valuable precursors for fine chemicals and pharmaceutics. However, most ADHs favor linear aliphatic and aromatic carbonyl compounds, and suitable biocatalysts with preference for cyclic ketones and diketones are still scarce. Among the few candidates, the alcohol dehydrogenase from Thauera aromatica (ThaADH) stands out with a high activity for the reduction of the cyclic α-diketone 1,2-cyclohexanedione to the corresponding α-hydroxy ketone. This study elucidates catalytic and structural features of the enzyme. ThaADH showed a remarkable thermal and pH stability as well as stability in the presence of polar solvents. A thorough description of the substrate scope combined with the resolution and description of the crystal structure, demonstrated a strong preference of ThaADH for cyclic α-substituted cyclohexanones, and indicated structural determinants responsible for the unique substrate acceptance.
- Professur für Molekulare Biotechnologie, Technische Universität Dresden, 01062, Dresden, Germany.
Organizational Affiliation: