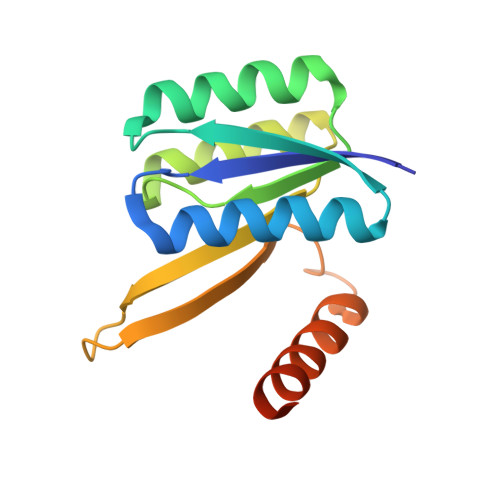
Structures of permuted halves of a modern ribose-binding protein.
Michel, F., Shanmugaratnam, S., Romero-Romero, S., Hocker, B.(2023) Acta Crystallogr D Struct Biol 79: 40-49
- PubMed: 36601806 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S205979832201186X
- Primary Citation Related Structures:
7QSP, 7QSQ - PubMed Abstract:
Periplasmic binding proteins (PBPs) are a class of proteins that participate in the cellular transport of various ligands. They have been used as model systems to study mechanisms in protein evolution, such as duplication, recombination and domain swapping. It has been suggested that PBPs evolved from precursors half their size. Here, the crystal structures of two permuted halves of a modern ribose-binding protein (RBP) from Thermotoga maritima are reported. The overexpressed proteins are well folded and show a monomer-dimer equilibrium in solution. Their crystal structures show partially noncanonical PBP-like fold type I conformations with structural deviations from modern RBPs. One of the half variants forms a dimer via segment swapping, suggesting a high degree of malleability. The structural findings on these permuted halves support the evolutionary hypothesis that PBPs arose via a duplication event of a flavodoxin-like protein and further support a domain-swapping step that might have occurred during the evolution of the PBP-like fold, a process that is necessary to generate the characteristic motion of PBPs essential to perform their functions.
- Department of Biochemistry, University of Bayreuth, 95447 Bayreuth, Germany.
Organizational Affiliation: