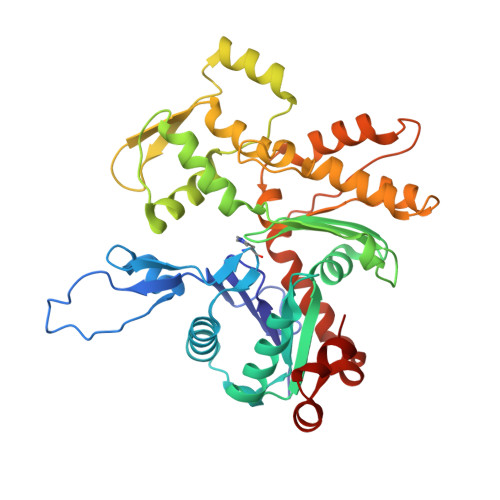
Structures from intact myofibrils reveal mechanism of thin filament regulation through nebulin.
Wang, Z., Grange, M., Pospich, S., Wagner, T., Kho, A.L., Gautel, M., Raunser, S.(2022) Science 375: eabn1934-eabn1934
- PubMed: 35175800 Search on PubMed
- DOI: https://doi.org/10.1126/science.abn1934
- Primary Citation Related Structures:
7QIM, 7QIN, 7QIO - PubMed Abstract:
In skeletal muscle, nebulin stabilizes and regulates the length of thin filaments, but the underlying mechanism remains nebulous. In this work, we used cryo-electron tomography and subtomogram averaging to reveal structures of native nebulin bound to thin filaments within intact sarcomeres. This in situ reconstruction provided high-resolution details of the interaction between nebulin and actin, demonstrating the stabilizing role of nebulin. Myosin bound to the thin filaments exhibited different conformations of the neck domain, highlighting its inherent structural variability in muscle. Unexpectedly, nebulin did not interact with myosin or tropomyosin, but it did interact with a troponin T linker through two potential binding motifs on nebulin, explaining its regulatory role. Our structures support the role of nebulin as a thin filament "molecular ruler" and provide a molecular basis for studying nemaline myopathies.
- Department of Structural Biochemistry, Max Planck Institute of Molecular Physiology, 44227 Dortmund, Germany.
Organizational Affiliation: