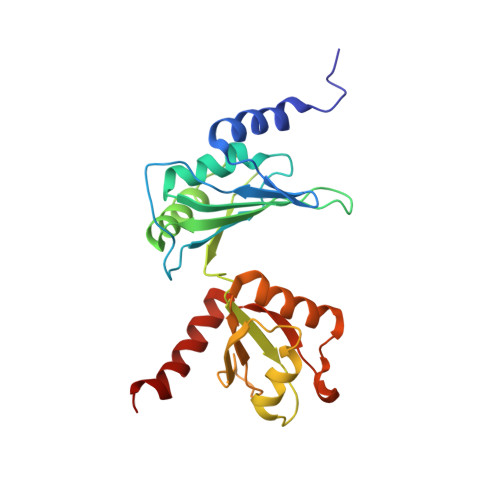
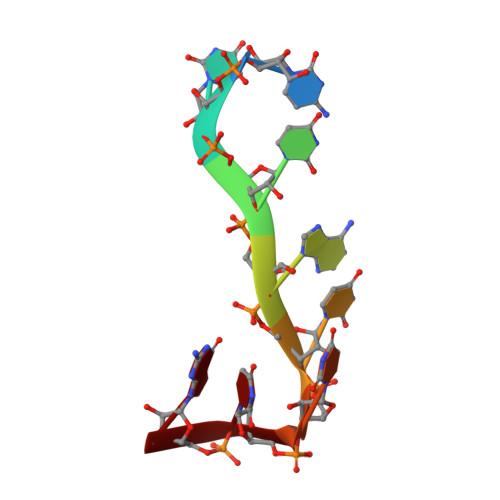
The solution structure of Dead End bound to AU-rich RNA reveals an unusual mode of tandem RRM-RNA recognition required for mRNA regulation.
Duszczyk, M.M., Wischnewski, H., Kazeeva, T., Arora, R., Loughlin, F.E., von Schroetter, C., Pradere, U., Hall, J., Ciaudo, C., Allain, F.H.(2022) Nat Commun 13: 5892-5892
- PubMed: 36202814 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33552-x
- Primary Citation Related Structures:
7Q4L - PubMed Abstract:
Dead End (DND1) is an RNA-binding protein essential for germline development through its role in post-transcriptional gene regulation. The molecular mechanisms behind selection and regulation of its targets are unknown. Here, we present the solution structure of DND1's tandem RNA Recognition Motifs (RRMs) bound to AU-rich RNA. The structure reveals how an NYAYUNN element is specifically recognized, reconciling seemingly contradictory sequence motifs discovered in recent genome-wide studies. RRM1 acts as a main binding platform, including atypical extensions to the canonical RRM fold. RRM2 acts cooperatively with RRM1, capping the RNA using an unusual binding pocket, leading to an unusual mode of tandem RRM-RNA recognition. We show that the consensus motif is sufficient to mediate upregulation of a reporter gene in human cells and that this process depends not only on RNA binding by the RRMs, but also on DND1's double-stranded RNA binding domain (dsRBD), which is dispensable for binding of a subset of targets in cellulo. Our results point to a model where DND1 target selection is mediated by a non-canonical mode of AU-rich RNA recognition by the tandem RRMs and a role for the dsRBD in the recruitment of effector complexes responsible for target regulation.
- Institute of Molecular Biology and Biophysics, ETH Zürich, 8093, Zürich, Switzerland. m.m.duszczyk@gmail.com.
Organizational Affiliation: