Synthesis of thieno[2,3-c]pyridine derived GRK2 inhibitors
Balo, T., Sapi, A., Kiss, A., Raimbaud, E., Paysant, J., Cattin, M.E., Berger, S., Kotschy, A., Faucher, N.(2022) Monatsh Chem
Experimental Data Snapshot
(2022) Monatsh Chem
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-adrenergic receptor kinase 1 | 697 | Homo sapiens | Mutation(s): 1 Gene Names: GRK2, ADRBK1, BARK, BARK1 EC: 2.7.11.15 | 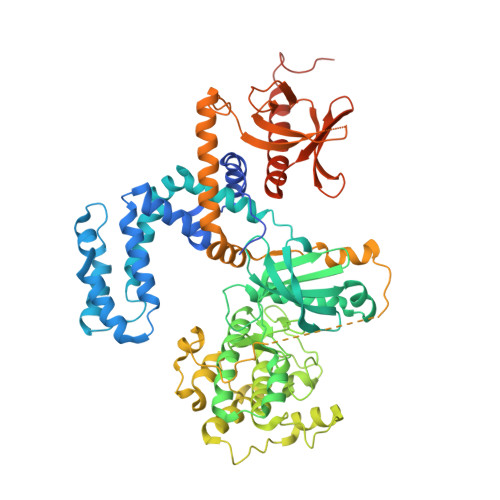 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P25098 GTEx: ENSG00000173020 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P25098 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 | 340 | Bos taurus | Mutation(s): 0 Gene Names: GNB1 | 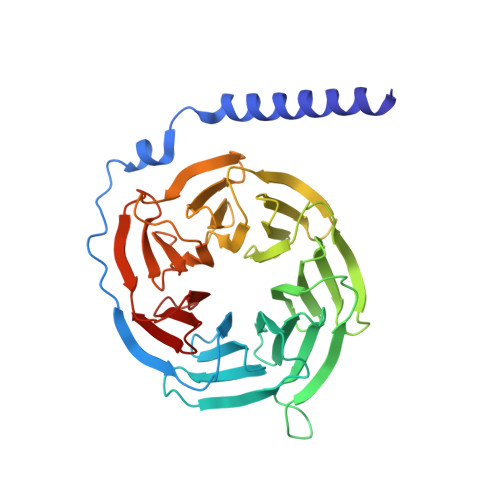 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62871 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 | C [auth G] | 77 | Bos taurus | Mutation(s): 1 Gene Names: GNG2 | 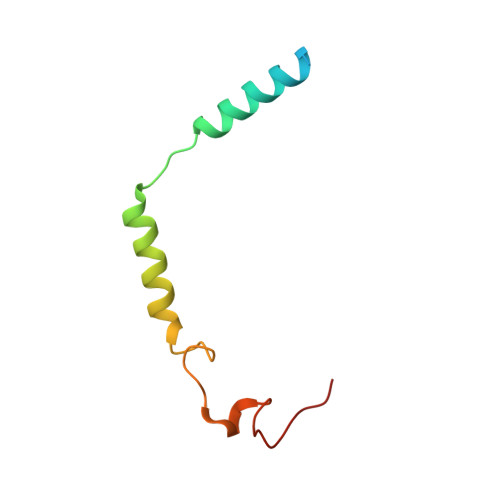 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63212 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 8DS (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | 4-chloranyl-N-[2-(4-chlorophenyl)ethyl]thieno[2,3-c]pyridine-2-carboxamide C16 H12 Cl2 N2 O S QRJZINDCUNBUGT-UHFFFAOYSA-N |  | ||
| 1PE Download:Ideal Coordinates CCD File | F [auth A] | PENTAETHYLENE GLYCOL C10 H22 O6 JLFNLZLINWHATN-UHFFFAOYSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | H [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | E [auth A], G [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 185.6 | α = 90 |
| b = 74.549 | β = 115.31 |
| c = 123.411 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| PHASER | phasing |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |