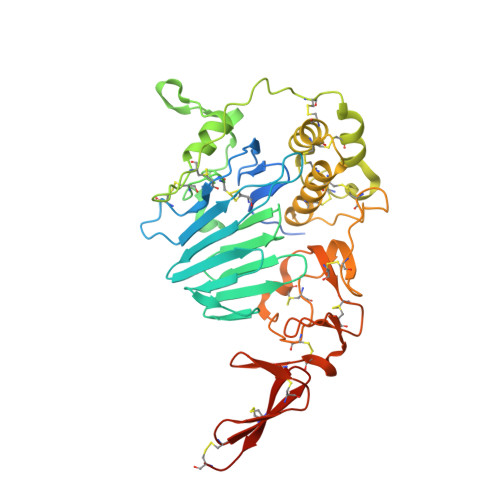
Intestinal mucin is a chaperone of multivalent copper.
Reznik, N., Gallo, A.D., Rush, K.W., Javitt, G., Fridmann-Sirkis, Y., Ilani, T., Nairner, N.A., Fishilevich, S., Gokhman, D., Chacon, K.N., Franz, K.J., Fass, D.(2022) Cell 185: 4206
- PubMed: 36206754 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2022.09.021
- Primary Citation Related Structures:
7PRL - PubMed Abstract:
Mucus protects the epithelial cells of the digestive and respiratory tracts from pathogens and other hazards. Progress in determining the molecular mechanisms of mucus barrier function has been limited by the lack of high-resolution structural information on mucins, the giant, secreted, gel-forming glycoproteins that are the major constituents of mucus. Here, we report how mucin structures we determined enabled the discovery of an unanticipated protective role of mucus: managing the toxic transition metal copper. Using two juxtaposed copper binding sites, one for Cu 2+ and the other for Cu 1+ , the intestinal mucin, MUC2, prevents copper toxicity by blocking futile redox cycling and the squandering of dietary antioxidants, while nevertheless permitting uptake of this important trace metal into cells. These findings emphasize the value of molecular structure in advancing mucosal biology, while introducing mucins, produced in massive quantities to guard extensive mucosal surfaces, as extracellular copper chaperones.
- Department of Chemical and Structural Biology, Weizmann Institute of Science, Rehovot 7610001, Israel.
Organizational Affiliation: