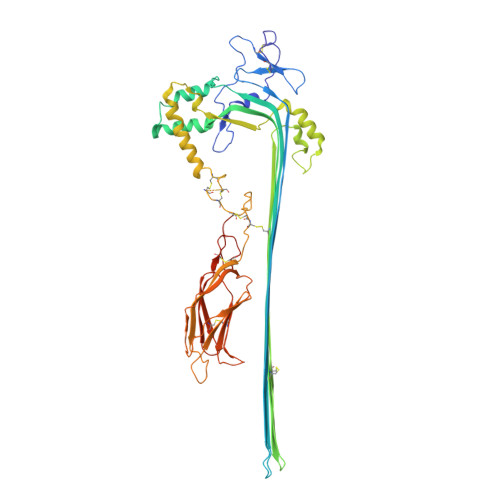
The pore conformation of lymphocyte perforin.
Ivanova, M.E., Lukoyanova, N., Malhotra, S., Topf, M., Trapani, J.A., Voskoboinik, I., Saibil, H.R.(2022) Sci Adv 8: eabk3147-eabk3147
- PubMed: 35148176 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abk3147
- Primary Citation Related Structures:
7PAG - PubMed Abstract:
Perforin is a pore-forming protein that facilitates rapid killing of pathogen-infected or cancerous cells by the immune system. Perforin is released from cytotoxic lymphocytes, together with proapoptotic granzymes, to bind to a target cell membrane where it oligomerizes and forms pores. The pores allow granzyme entry, which rapidly triggers the apoptotic death of the target cell. Here, we present a 4-Å resolution cryo-electron microscopy structure of the perforin pore, revealing previously unidentified inter- and intramolecular interactions stabilizing the assembly. During pore formation, the helix-turn-helix motif moves away from the bend in the central β sheet to form an intermolecular contact. Cryo-electron tomography shows that prepores form on the membrane surface with minimal conformational changes. Our findings suggest the sequence of conformational changes underlying oligomerization and membrane insertion, and explain how several pathogenic mutations affect function.
- Institute of Structural and Molecular Biology, Birkbeck, University of London, Malet St, London WC1E 7HX, UK.
Organizational Affiliation: