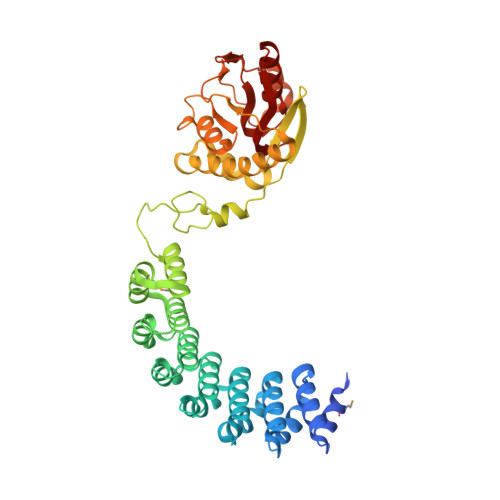
ComFC mediates transport and handling of single-stranded DNA during natural transformation.
Damke, P.P., Celma, L., Kondekar, S.M., Di Guilmi, A.M., Marsin, S., Depagne, J., Veaute, X., Legrand, P., Walbott, H., Vercruyssen, J., Guerois, R., Quevillon-Cheruel, S., Radicella, J.P.(2022) Nat Commun 13: 1961-1961
- PubMed: 35414142 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-29494-z
- Primary Citation Related Structures:
7P0H - PubMed Abstract:
The ComFC protein is essential for natural transformation, a process that plays a major role in the spread of antibiotic resistance genes and virulence factors across bacteria. However, its role remains largely unknown. Here, we show that Helicobacter pylori ComFC is involved in DNA transport through the cell membrane, and is required for the handling of the single-stranded DNA once it is delivered into the cytoplasm. The crystal structure of ComFC includes a zinc-finger motif and a putative phosphoribosyl transferase domain, both necessary for the protein's in vivo activity. Furthermore, we show that ComFC is a membrane-associated protein with affinity for single-stranded DNA. Our results suggest that ComFC provides the link between the transport of the transforming DNA into the cytoplasm and its handling by the recombination machinery.
- Université Paris-Saclay, CEA, Stabilité Génétique Cellules Souches et Radiations, Institut de Biologie François Jacob, F-92260, Fontenay aux Roses, France.
Organizational Affiliation: